| DPOGS200036 | ||
|---|---|---|
| Transcript | DPOGS200036-TA | 396 bp |
| Protein | DPOGS200036-PA | 131 aa |
| Genomic position | DPSCF300538 - 33416-35712 | |
| RNAseq coverage | 62x (Rank: top 68%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL013881 | 2e-41 | 64.62% | |
| Bombyx | BGIBMGA010237-TA | 5e-34 | 56.10% | |
| Drosophila | Gs1l-PB | 2e-30 | 55.37% | |
| EBI UniRef50 | UniRef50_E0VBZ4 | 9e-31 | 54.17% | 2-deoxyglucose-6-phosphate phosphatase, putative n=3 Tax=Coelomata RepID=E0VBZ4_PEDHC |
| NCBI RefSeq | XP_002052751.1 | 1e-32 | 57.98% | GJ17733 [Drosophila virilis] |
| NCBI nr blastp | gi|195388160 | 3e-31 | 57.98% | GJ17733 [Drosophila virilis] |
| NCBI nr blastx | gi|195388160 | 4e-30 | 57.98% | GJ17733 [Drosophila virilis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016787 | 1.3e-12 | hydrolase activity | |
| KEGG pathway | ath:AT4G21470 | 1e-11 | ||
| K00861 (RFK, FMN1) | maps-> | Riboflavin metabolism | ||
| InterPro domain | [1-120] IPR023214 | 5.5e-25 | HAD-like domain | |
| [18-86] IPR006402 | 1.3e-12 | HAD-superfamily hydrolase, subfamily IA, variant 3 | ||
| Orthology group | ||||
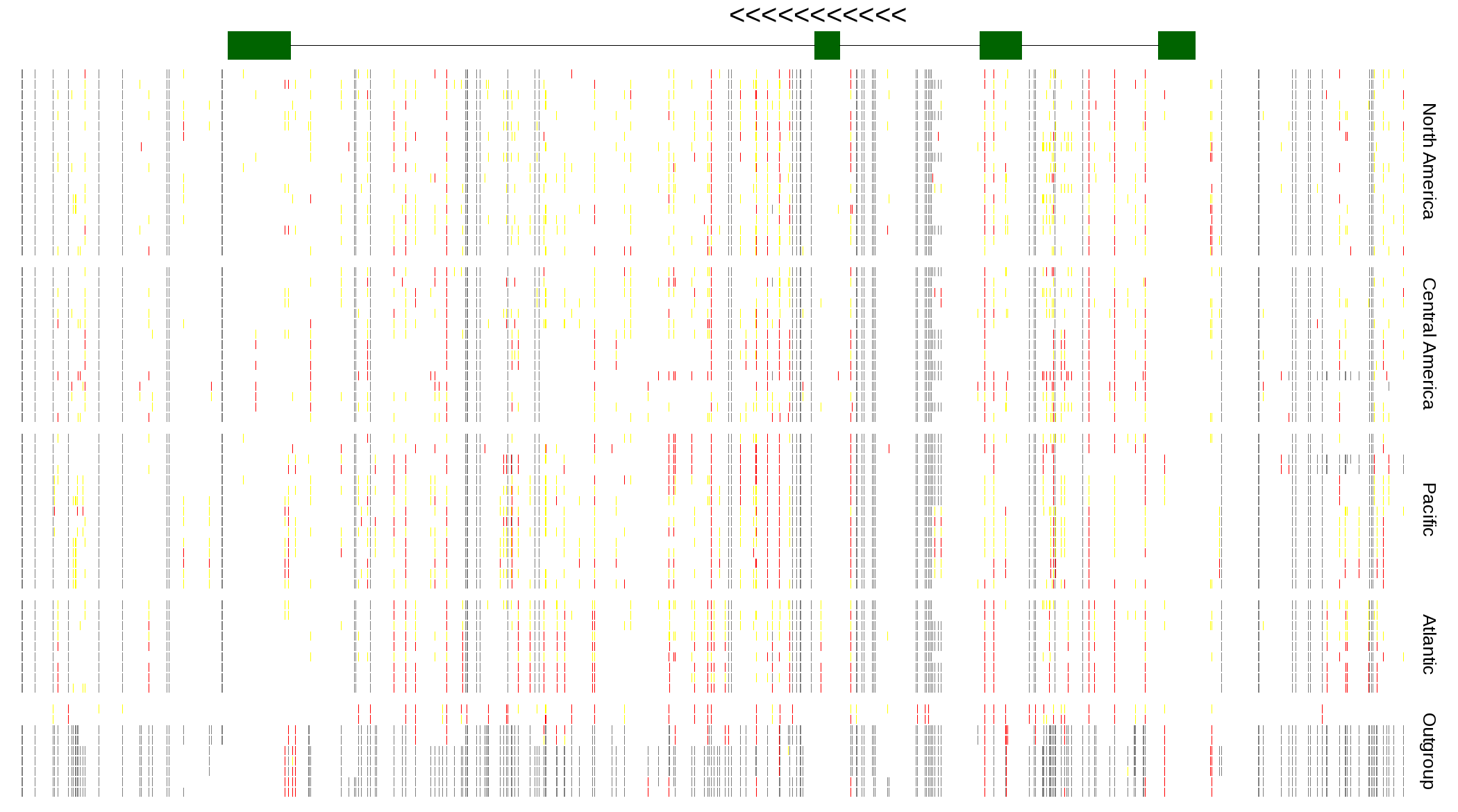
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS200036-TA
ATGGCGTTAGCAACTAACTCCACTGCCCAGGCCGTCCGACTCCACGCTACGGCTCGACCGAAGCTATTTGGACTGTTCCACCATAAAGTAAGCGTGACGGATCCGGAAGTCCTTCGCGGTAAGCCCTACCCTGATATATACATGGTAGCTGCAGCAAGATTTCCAGAGAAACCCAAGGCGAAACAGTGCCTCGTATTTGAAGATTCCTTTGTTGGTGTGAAGTCAGCAGTCGAAGCGGGAATGCAGGTGGTCATGATACCAGACAGTCGTATCGATCGTGAGCAAACCCGACAAGCAACGTTAGTCATACGGTCCCTTCTAGACTTCCAACCGGAACTCTTCGGACTACCGCCGTTTGATGACACACCGAGACCTTGTTCTAGAATTGAAAAGTAA
>DPOGS200036-PA
MALATNSTAQAVRLHATARPKLFGLFHHKVSVTDPEVLRGKPYPDIYMVAAARFPEKPKAKQCLVFEDSFVGVKSAVEAGMQVVMIPDSRIDREQTRQATLVIRSLLDFQPELFGLPPFDDTPRPCSRIEK-