| DPOGS200166 | ||
|---|---|---|
| Transcript | DPOGS200166-TA | 567 bp |
| Protein | DPOGS200166-PA | 188 aa |
| Genomic position | DPSCF300128 + 333954-335223 | |
| RNAseq coverage | 1766x (Rank: top 7%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006881 | 2e-94 | 96.43% | |
| Bombyx | BGIBMGA002918-TA | 2e-95 | 97.04% | |
| Drosophila | UbcD2-PB | 8e-92 | 81.09% | |
| EBI UniRef50 | UniRef50_Q96LR5 | 2e-82 | 87.27% | Ubiquitin-conjugating enzyme E2 E2 n=57 Tax=Coelomata RepID=UB2E2_HUMAN |
| NCBI RefSeq | XP_971173.1 | 7e-94 | 95.86% | PREDICTED: similar to AGAP007952-PA [Tribolium castaneum] |
| NCBI nr blastp | gi|91090117 | 1e-92 | 95.86% | PREDICTED: similar to AGAP007952-PA [Tribolium castaneum] |
| NCBI nr blastx | gi|332028072 | 3e-98 | 92.97% | Ubiquitin-conjugating enzyme E2-24 kDa [Acromyrmex echinatior] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016881 | 2.6e-46 | acid-amino acid ligase activity | |
| KEGG pathway | tca:659808 | 2e-93 | ||
| K06689 (UBE2D_E, UBC4, UBC5) | maps-> | Ubiquitin mediated proteolysis | ||
| Protein processing in endoplasmic reticulum | ||||
| InterPro domain | [31-187] IPR016135 | 1.8e-73 | Ubiquitin-conjugating enzyme/RWD-like | |
| [46-181] IPR000608 | 2.6e-46 | Ubiquitin-conjugating enzyme, E2 | ||
| Orthology group | MCL11092 | Multiple-copy universal gene | ||
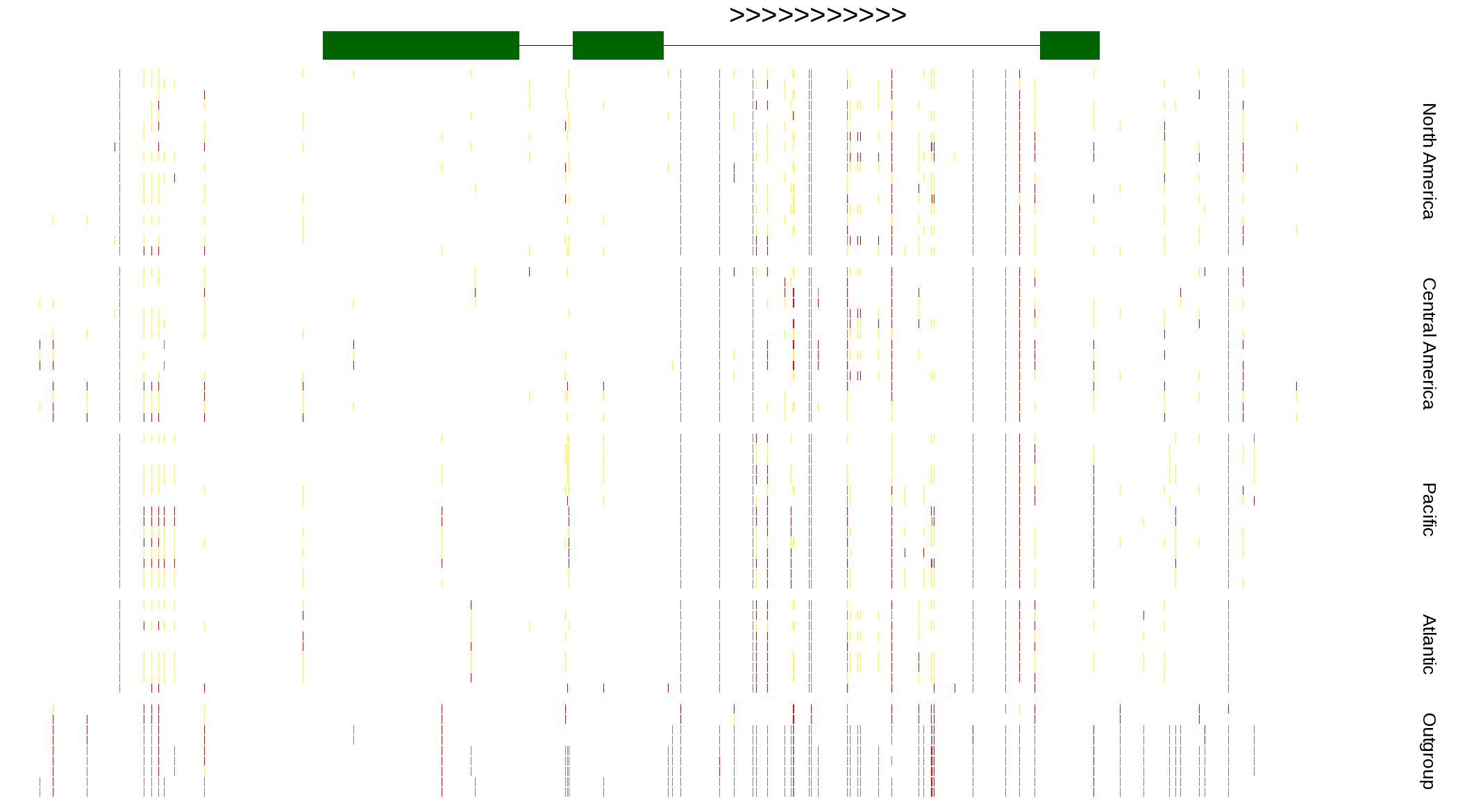
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS200166-TA
ATGTCTGGTGCGTCTGGTTCATCAGGGAGTGGTACAGGGCGAGGCAGAGGTGGTAACGCAGTCGGGGATAAGCCGGATACTAAAGAAAATAAACCTAACGCTAAAATGTCGAAAGCACTTGGTACATCAGCCAAGAGAATCCAAAAGGAGCTGGCCGAGATCACACTAGATCCCCCGCCTAACTGTAGTGCAGGTCCTAAGGGTGACAATTTATATGAATGGGTCTCAACAATATTAGGGCCGCCCGGCTCGGTTTATGAAGGTGGCGTCTTCTTTTTAGATATTCATTTTTCACCGGAATATCCCTTCAAACCTCCTAAGGTGACGTTTCGTACGAGGATATACCACTGTAATATAAACAGCCAGGGCGTTATATGTTTAGATATTTTGAAAGACAATTGGTCGCCGGCACTCACCATTTCAAAAGTACTATTATCTATATGCTCCCTCTTGACTGACTGTAATCCTGCCGATCCCCTCGTAGGCAGCATTGCGACGCAGTACCTCCAGAACAGAGAGGAACACGATCGTATCGCCCGTCTGTGGACCAAGCGCTATGCTACATGA
>DPOGS200166-PA
MSGASGSSGSGTGRGRGGNAVGDKPDTKENKPNAKMSKALGTSAKRIQKELAEITLDPPPNCSAGPKGDNLYEWVSTILGPPGSVYEGGVFFLDIHFSPEYPFKPPKVTFRTRIYHCNINSQGVICLDILKDNWSPALTISKVLLSICSLLTDCNPADPLVGSIATQYLQNREEHDRIARLWTKRYAT-