| DPOGS200209 | ||
|---|---|---|
| Transcript | DPOGS200209-TA | 639 bp |
| Protein | DPOGS200209-PA | 212 aa |
| Genomic position | DPSCF300093 + 489772-492781 | |
| RNAseq coverage | 0x (Rank: top 95%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | % | |||
| Bombyx | % | |||
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_D7F179 | 4e-40 | 60.14% | Endonuclease-reverse transcriptase n=3 Tax=Obtectomera RepID=D7F179_BOMMO |
| NCBI RefSeq | XP_001187905.1 | 2e-23 | 45.16% | PREDICTED: similar to protein F28E10.3 [imported] - Caenorhabditis elegans [Strongylocentrotus purpuratus] |
| NCBI nr blastp | gi|298204367 | 1e-39 | 60.14% | endonuclease-reverse transcriptase [Bombyx mori] |
| NCBI nr blastx | gi|298204367 | 9e-38 | 60.58% | endonuclease-reverse transcriptase [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003964 | 8.4e-12 | RNA-directed DNA polymerase activity | |
| GO:0003723 | 8.4e-12 | RNA binding | ||
| GO:0006278 | 8.4e-12 | RNA-dependent DNA replication | ||
| KEGG pathway | ||||
| InterPro domain | [75-199] IPR000477 | 8.4e-12 | Reverse transcriptase | |
| Orthology group | MCL27789 | Specific divergent | ||
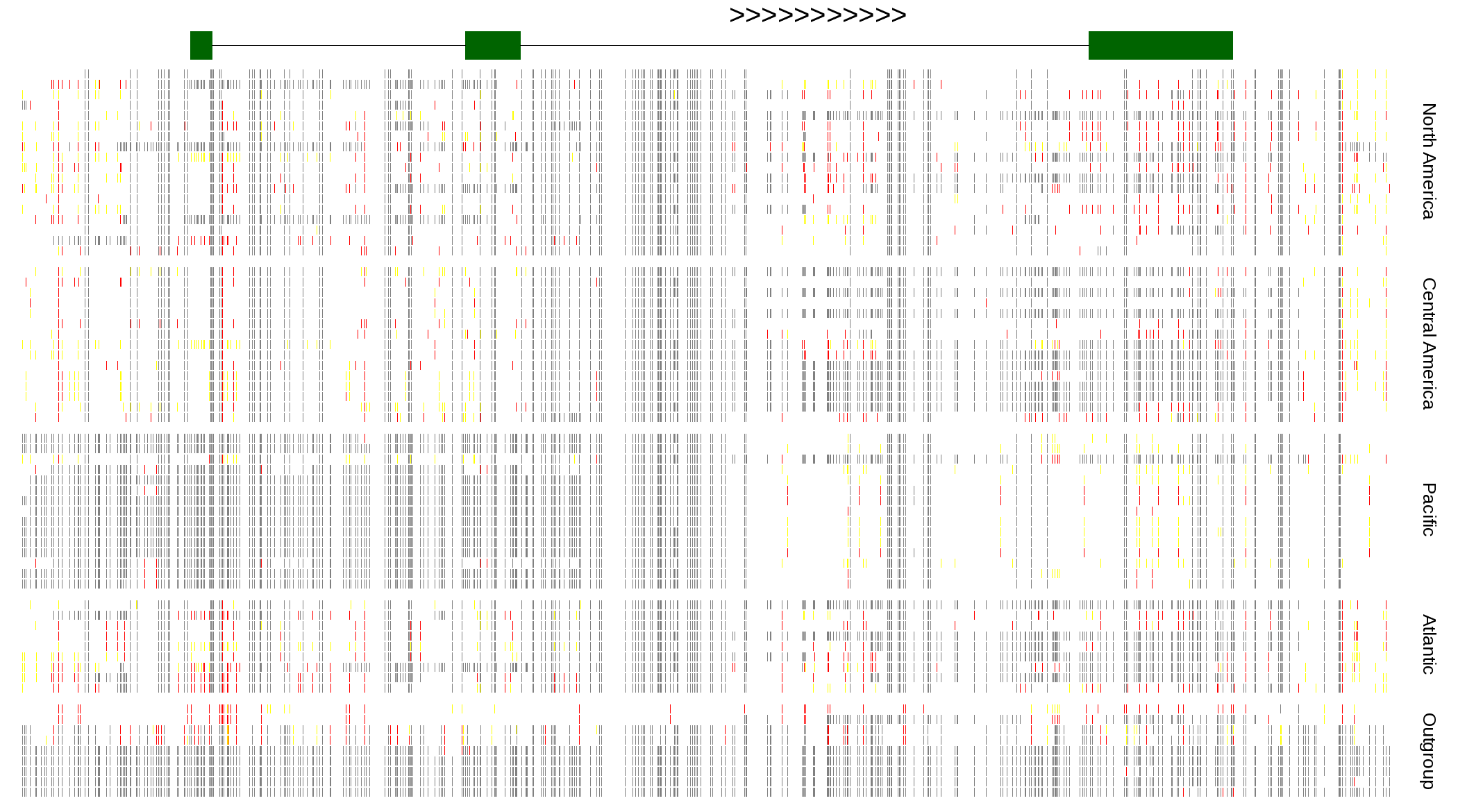
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS200209-TA
ATGAACTTTAAAGTCAAAGACGTTGAAGCCATACCTGAAATTTTAGGATCGGAAATGGAACAAGTAGACAGAAGCAGTGAAGTAGGTCTAGAAATAAACACTAGCAATACTAAAGTGATGACCAACTCAACACCCACAGACATAACAGTCAACAGACAGAAACTCGAATATGTATACCTAGGCCAGATAATATCGCAAAAGGACCAAATGTCCATAGAAATAGCGAAAATAAAACTAGAATCTACAGGAGATGCTTTCACAATTAAAAGGGGAGACAGGCAAGGTGACCCACTCTCACCAAAGCTGTTTACAGCCCTACTAGAACAAATGTTTAGAAAACTGAATTGGGAAAATCAAGGGCTAAACATAAATGGTGCACGATTGAGCCATCAAAGAGTCGCAGACGACCTTGTCATATTAGAATCGAATCCACTAACACTAGAATCAATGATACAAACCCTAGTAGACAGAAGCAGTGAAGTATGTCTAGAAATAATCACTAGCAAAACTAAAGGGATGACCAACTCAACACCCACAGACATAACAATCAACAGACAGAAACTCGAATATGTAGAGGAATATGTTTACCTAGGCCACATAATATTGCAAAAAGACCAAATGTCCATAGAAATAGTTTAA
>DPOGS200209-PA
MNFKVKDVEAIPEILGSEMEQVDRSSEVGLEINTSNTKVMTNSTPTDITVNRQKLEYVYLGQIISQKDQMSIEIAKIKLESTGDAFTIKRGDRQGDPLSPKLFTALLEQMFRKLNWENQGLNINGARLSHQRVADDLVILESNPLTLESMIQTLVDRSSEVCLEIITSKTKGMTNSTPTDITINRQKLEYVEEYVYLGHIILQKDQMSIEIV-