| DPOGS200593 | ||
|---|---|---|
| Transcript | DPOGS200593-TA | 456 bp |
| Protein | DPOGS200593-PA | 151 aa |
| Genomic position | DPSCF300076 - 667751-669081 | |
| RNAseq coverage | 15112x (Rank: top 1%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL001035 | 1e-84 | 99.34% | |
| Bombyx | BGIBMGA011282-TA | 2e-84 | 99.34% | |
| Drosophila | RpS13-PB | 1e-76 | 88.74% | |
| EBI UniRef50 | UniRef50_P62277 | 3e-76 | 90.73% | 40S ribosomal protein S13 n=441 Tax=root RepID=RS13_HUMAN |
| NCBI RefSeq | NP_001091754.1 | 4e-83 | 99.34% | ribosomal protein S13 [Bombyx mori] |
| NCBI nr blastp | gi|54039367 | 2e-82 | 100.00% | ribosomal protein S13 [Spodoptera frugiperda] |
| NCBI nr blastx | gi|54039367 | 4e-80 | 100.00% | ribosomal protein S13 [Spodoptera frugiperda] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005840 | 1.2e-28 | ribosome | |
| GO:0006412 | 1.2e-28 | translation | ||
| GO:0003735 | 1.2e-28 | structural constituent of ribosome | ||
| GO:0005622 | 5.9e-24 | intracellular | ||
| KEGG pathway | phu:Phum_PHUM357760 | 3e-77 | ||
| K02953 (RP-S13e, RPS13) | maps-> | Ribosome | ||
| InterPro domain | [1-60] IPR012606 | 1.2e-28 | Ribosomal protein S13/S15, N-terminal | |
| [66-150] IPR000589 | 5.9e-24 | Ribosomal protein S15 | ||
| [65-142] IPR009068 | 3.7e-22 | S15/NS1, RNA-binding | ||
| Orthology group | MCL11047 | Multiple-copy universal gene | ||
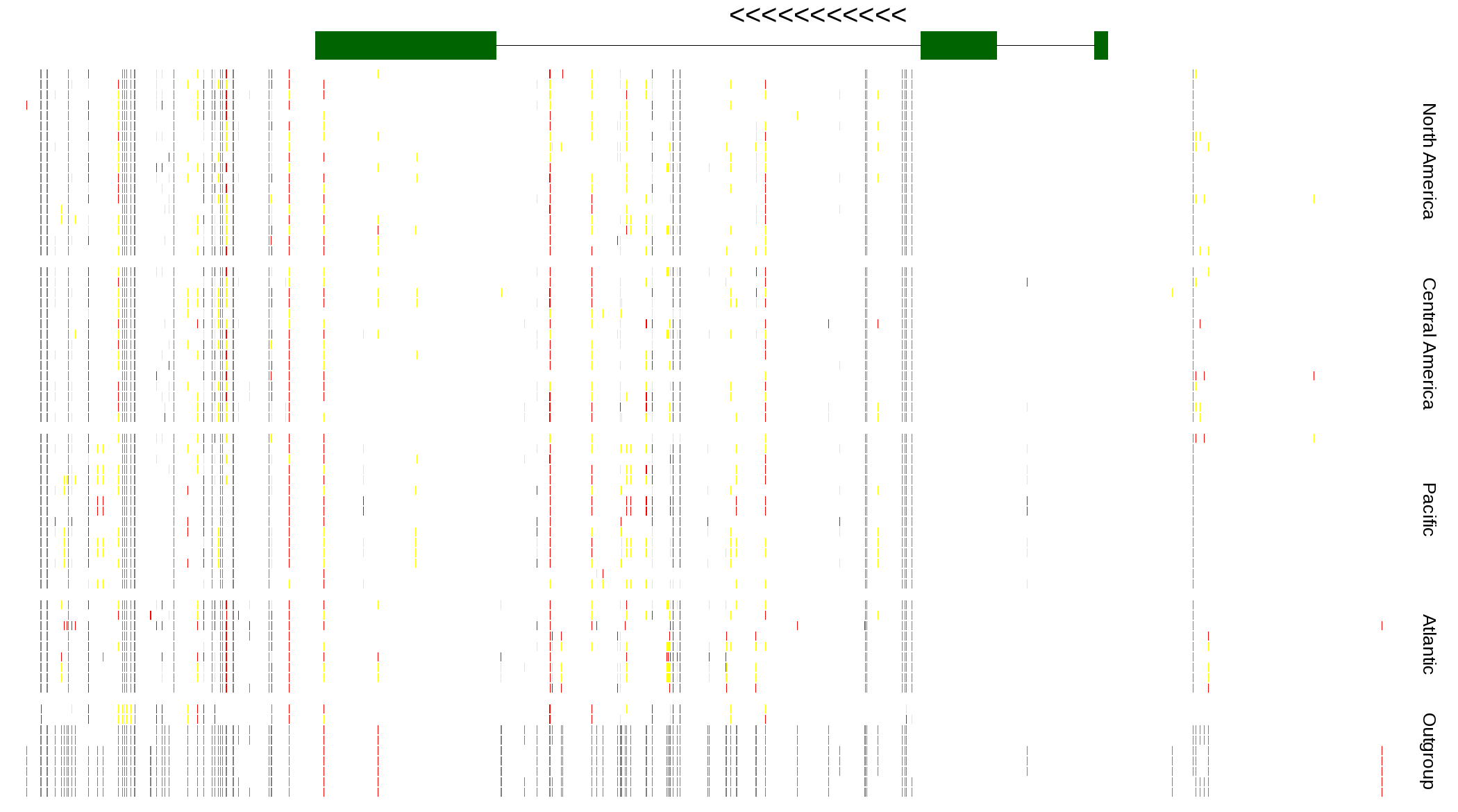
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS200593-TA
ATGGGTCGTATGCACGCACCTGGGAAAGGTATCTCACAGTCGGCGTTACCATACCGCCGCAGTGTCCCCACCTGGTTAAAACTCACAGCTGATGATGTTAAGGAACAGATTTTCAAACTCGGCAAGAAGGGTCTTACTCCATCCCAAATCGGTGTCATGCTCAGGGATTCCCACGGAGTTGCCCAAGTCAGATTTGTTACCGGCAAAAAGATTCTCCGTATCATGAAAGCCATGGGTCTAGCTCCGGATCTACCTGAAGACTTGTACTACCTGATCAAGAAAGCCGTCGCTATGAGGAAGCACTTGGAACGTAACAGGAAAGACAAAGACAGCAAATTCAGGCTCATTCTCGTAGAGTCTAGAATCCACAGGCTCGCACGCTACTACAAAACCAAGAGCGTGCTGCCCCCTAACTGGAAGTATGAGTCGAGCACCGCCTCTGCTCTTGTGGCTTAA
>DPOGS200593-PA
MGRMHAPGKGISQSALPYRRSVPTWLKLTADDVKEQIFKLGKKGLTPSQIGVMLRDSHGVAQVRFVTGKKILRIMKAMGLAPDLPEDLYYLIKKAVAMRKHLERNRKDKDSKFRLILVESRIHRLARYYKTKSVLPPNWKYESSTASALVA-