| DPOGS200760 | ||
|---|---|---|
| Transcript | DPOGS200760-TA | 252 bp |
| Protein | DPOGS200760-PA | 83 aa |
| Genomic position | DPSCF300556 - 280-8812 | |
| RNAseq coverage | 213x (Rank: top 46%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL005536 | 7e-37 | 85.37% | |
| Bombyx | BGIBMGA004224-TA | 3e-18 | 78.43% | |
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_H0YNJ6 | 3e-24 | 72.60% | Guanosine monophosphate reductase 2 n=6 Tax=Catarrhini RepID=H0YNJ6_HUMAN |
| NCBI RefSeq | XP_001603068.1 | 4e-32 | 80.00% | PREDICTED: similar to Gmpr2 protein [Nasonia vitripennis] |
| NCBI nr blastp | gi|156547587 | 1e-30 | 80.00% | PREDICTED: GMP reductase 1-like [Nasonia vitripennis] |
| NCBI nr blastx | gi|156547587 | 3e-28 | 80.00% | PREDICTED: GMP reductase 1-like [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0008152 | 5.1e-21 | metabolic process | |
| GO:0003824 | 5.1e-21 | catalytic activity | ||
| GO:0055114 | 1.7e-13 | oxidation-reduction process | ||
| KEGG pathway | nvi:100119268 | 1e-31 | ||
| K00364 (E1.7.1.7, guaC) | maps-> | Purine metabolism | ||
| InterPro domain | [2-72] IPR013785 | 5.1e-21 | Aldolase-type TIM barrel | |
| [9-73] IPR001093 | 1.7e-13 | IMP dehydrogenase/GMP reductase | ||
| Orthology group | ||||
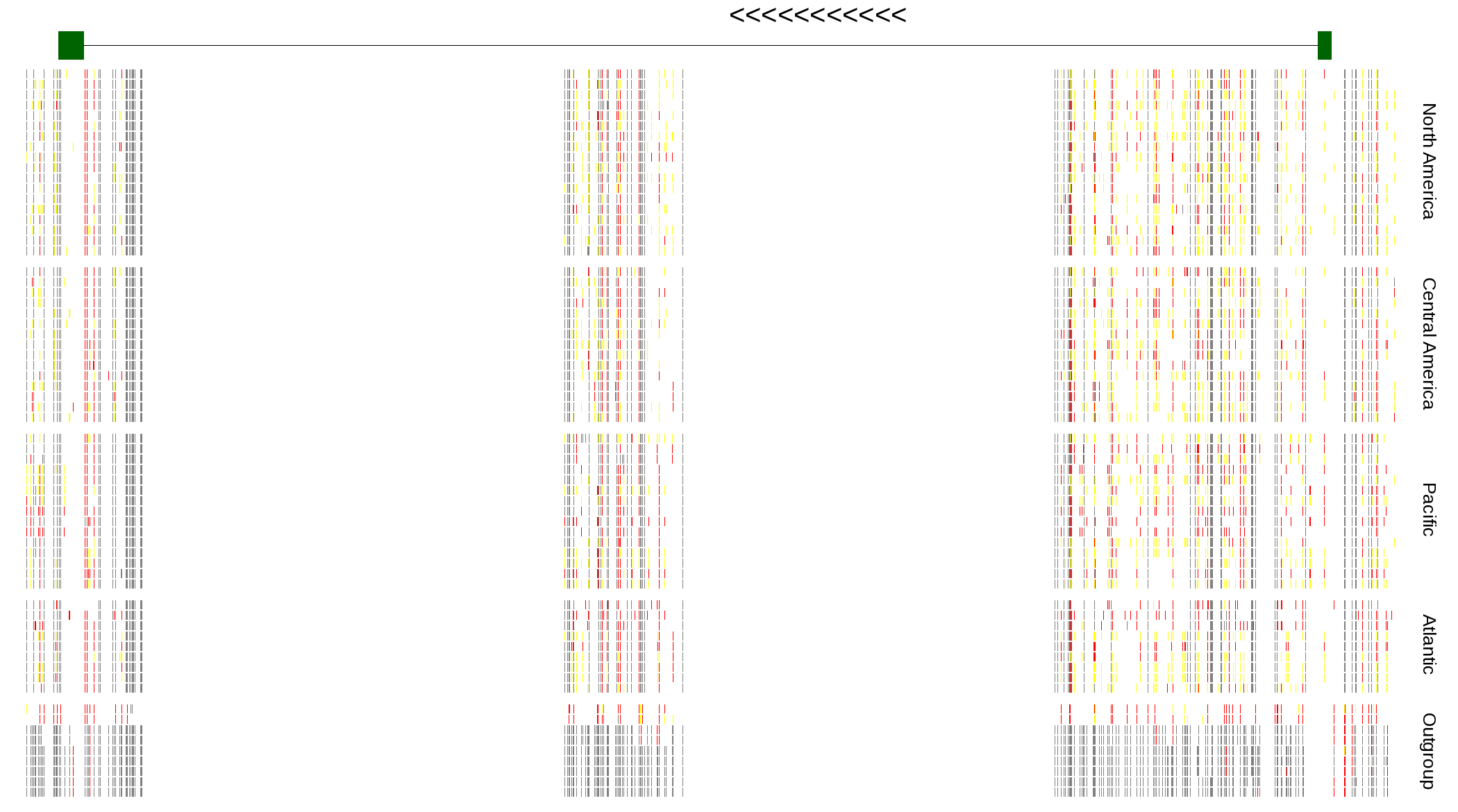
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS200760-TA
ATGCCTAATATTATAAATGAGGTTAAATTGGATTTTAAAGACGTGTTACTGCGGCCTAAACGGAGCACACTGAAAAGTAGAAACGATGTGGATCTGTTCAGGGAGATAACGTTCCGTAACTCTGGCCAGACTTACCGCGGCGTTCCCGTCATGGCGAGCAATATGGACACCGTGGGCACATTCGAAATGGCCATGGAATTGTCGAAGGTTGGTCGCTTTAGATTGGAATGGCATTGGTATCCGATGACATAA
>DPOGS200760-PA
MPNIINEVKLDFKDVLLRPKRSTLKSRNDVDLFREITFRNSGQTYRGVPVMASNMDTVGTFEMAMELSKVGRFRLEWHWYPMT-