| DPOGS200940 | ||
|---|---|---|
| Transcript | DPOGS200940-TA | 525 bp |
| Protein | DPOGS200940-PA | 174 aa |
| Genomic position | DPSCF300215 - 271053-272018 | |
| RNAseq coverage | 61x (Rank: top 68%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL004676 | 2e-27 | 66.29% | |
| Bombyx | BGIBMGA010205-TA | 2e-51 | 90.98% | |
| Drosophila | n-syb-PE | 9e-45 | 80.37% | |
| EBI UniRef50 | UniRef50_B4PDQ3 | 3e-44 | 68.99% | GE20941 n=11 Tax=Coelomata RepID=B4PDQ3_DROYA |
| NCBI RefSeq | XP_002432199.1 | 4e-47 | 75.40% | synaptobrevin, putative [Pediculus humanus corporis] |
| NCBI nr blastp | gi|242023558 | 5e-46 | 75.40% | synaptobrevin, putative [Pediculus humanus corporis] |
| NCBI nr blastx | gi|242023558 | 4e-45 | 82.46% | synaptobrevin, putative [Pediculus humanus corporis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016021 | 4e-31 | integral to membrane | |
| GO:0016192 | 4e-31 | vesicle-mediated transport | ||
| KEGG pathway | phu:Phum_PHUM576430 | 1e-46 | ||
| K13504 (VAMP2) | maps-> | Salivary secretion | ||
| SNARE interactions in vesicular transport | ||||
| Vasopressin-regulated water reabsorption | ||||
| InterPro domain | [58-139] IPR001388 | 4e-31 | Synaptobrevin | |
| Orthology group | MCL25714 | Lepidoptera specific | ||
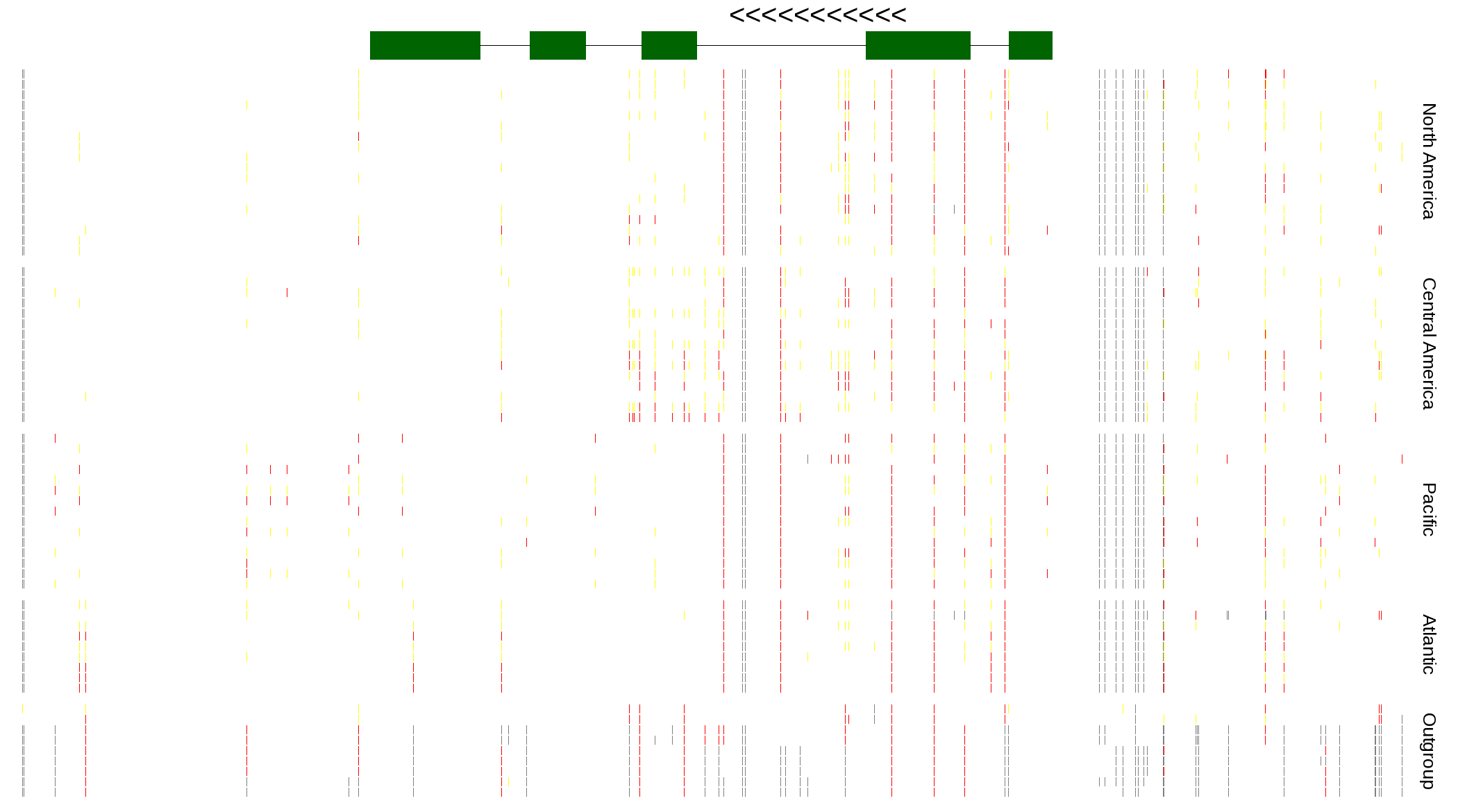
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS200940-TA
ATGTTAGCTTGGCGTCCTTGCGCTGCTGATATGGCATCGTGCCAGACTGGTATTGAAAAAAGGGCAGCACTAGAAGGCGGCACTCCTGGGGGTCTTGCCCCGGCGGAGCCCCAGACCGGTCCAGATGGTGAAATCATCGGAGGACCTCGTACTCCACAACAGATTGCTGCCCAAAGGAGGCTACAACAGACCCAGGCGCAAGTCGATGAAGTTGTGGATATTATGAAATCTAACGTCGAGAAGGTATTAGACAGAGATGCCAAACTATCAGAACTTGATGATAGAGCAGACGCGCTTCAGCAGGGAGCATCGCAGTTTGAACAGCAAGCTAAGAGTTTGAAGAACAAATTTTGGCTTCAAAATCTTAAGATGATGATTATTATGGGGGTGATTGGTATCGTTATCATTGGACTCCTGTTTGGTGAGTATACTAAGATTGTTACTATGAAACTAGATGTTACTTATGATTGTTTAGATGTGTCATGGCCGTTAGTACCTGTGATATTTGTTAAAATAGTGTTTTAA
>DPOGS200940-PA
MLAWRPCAADMASCQTGIEKRAALEGGTPGGLAPAEPQTGPDGEIIGGPRTPQQIAAQRRLQQTQAQVDEVVDIMKSNVEKVLDRDAKLSELDDRADALQQGASQFEQQAKSLKNKFWLQNLKMMIIMGVIGIVIIGLLFGEYTKIVTMKLDVTYDCLDVSWPLVPVIFVKIVF-