| DPOGS201505 | ||
|---|---|---|
| Transcript | DPOGS201505-TA | 384 bp |
| Protein | DPOGS201505-PA | 127 aa |
| Genomic position | DPSCF300006 + 951198-951780 | |
| RNAseq coverage | 10x (Rank: top 84%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL016991 | 6e-17 | 48.65% | |
| Bombyx | BGIBMGA006513-TA | 7e-18 | 55.22% | |
| Drosophila | CG10527-PA | 4e-16 | 45.95% | |
| EBI UniRef50 | UniRef50_E3WQH1 | 3e-15 | 39.18% | Putative uncharacterized protein n=1 Tax=Anopheles darlingi RepID=E3WQH1_ANODA |
| NCBI RefSeq | XP_001599775.1 | 4e-18 | 55.41% | PREDICTED: similar to farnesoic acid o-methyltransferase-like isoform 1 protein, partial [Nasonia vitripennis] |
| NCBI nr blastp | gi|183584866 | 4e-16 | 57.14% | putative farnesoic acid O-methyltransferase [Ceratitis capitata] |
| NCBI nr blastx | gi|345495861 | 2e-24 | 60.00% | PREDICTED: hypothetical protein LOC100114909, partial [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ||||
| InterPro domain | [57-127] IPR006616 | 2e-22 | DM9 repeat | |
| Orthology group | ||||
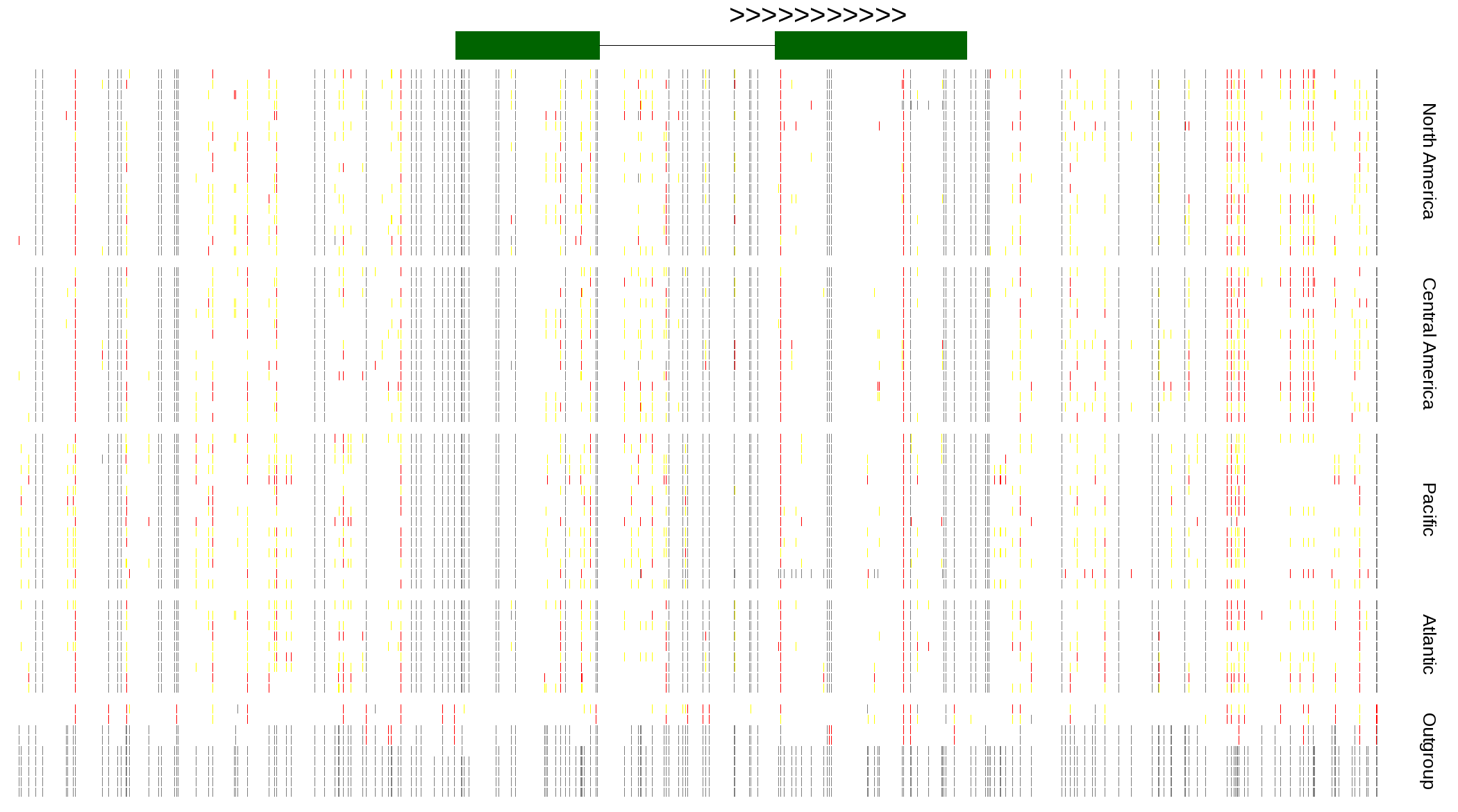
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS201505-TA
ATGGTAACGCGCCGGCAAATGCTGGGGAAACTGGGGAACCTATATACGTGGCCAGAGCTGAACATGAGGGAGACCTCGTTCCTGGAAAACTTGTATCCTCCCATAAAGGAACTTCCGTTCCTTAGGGTGTTAAGGAGATCTCTAAGAACGAATACGAGGTGCGAGAGTACTTACCAATGGGTACCCGCTCGAGGTGGTTCTGTGCCAGCTAATGCTCTTAAAGCAGGTATGACTAAGAGTGGGAAAACTTTATATATGGGCCGATTAAATCACCTCAACTCACTCACACCAGGAAAAGTACACCCCTCGCAAGGTGCTTATTATATACCCCTTGGTGGAGAAGAATTAAAATTTACCGACTATGAAGTCGTCTTGGCAAATTGA
>DPOGS201505-PA
MVTRRQMLGKLGNLYTWPELNMRETSFLENLYPPIKELPFLRVLRRSLRTNTRCESTYQWVPARGGSVPANALKAGMTKSGKTLYMGRLNHLNSLTPGKVHPSQGAYYIPLGGEELKFTDYEVVLAN-