| DPOGS201570 | ||
|---|---|---|
| Transcript | DPOGS201570-TA | 555 bp |
| Protein | DPOGS201570-PA | 184 aa |
| Genomic position | DPSCF300201 + 142935-145017 | |
| RNAseq coverage | 1x (Rank: top 93%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL010923 | 3e-62 | 63.16% | |
| Bombyx | BGIBMGA006118-TA | 3e-37 | 43.46% | |
| Drosophila | fs(2)ltoPP43-PA | 7e-06 | 23.60% | |
| EBI UniRef50 | UniRef50_UPI00021A779F | 8e-18 | 34.21% | UPI00021A779F related cluster n=3 Tax=unknown RepID=UPI00021A779F |
| NCBI RefSeq | XP_001122521.1 | 7e-23 | 38.92% | PREDICTED: similar to female sterile (2) ltoPP43 CG10528-PA, isoform A [Apis mellifera] |
| NCBI nr blastp | gi|383864526 | 3e-21 | 37.89% | PREDICTED: probable RNA polymerase II nuclear localization protein SLC7A6OS-like [Megachile rotundata] |
| NCBI nr blastx | gi|270007526 | 2e-24 | 32.29% | hypothetical protein TcasGA2_TC014121 [Tribolium castaneum] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ||||
| InterPro domain | [93-161] IPR013883 | 3.8e-06 | Transcription factor Iwr1 | |
| Orthology group | ||||
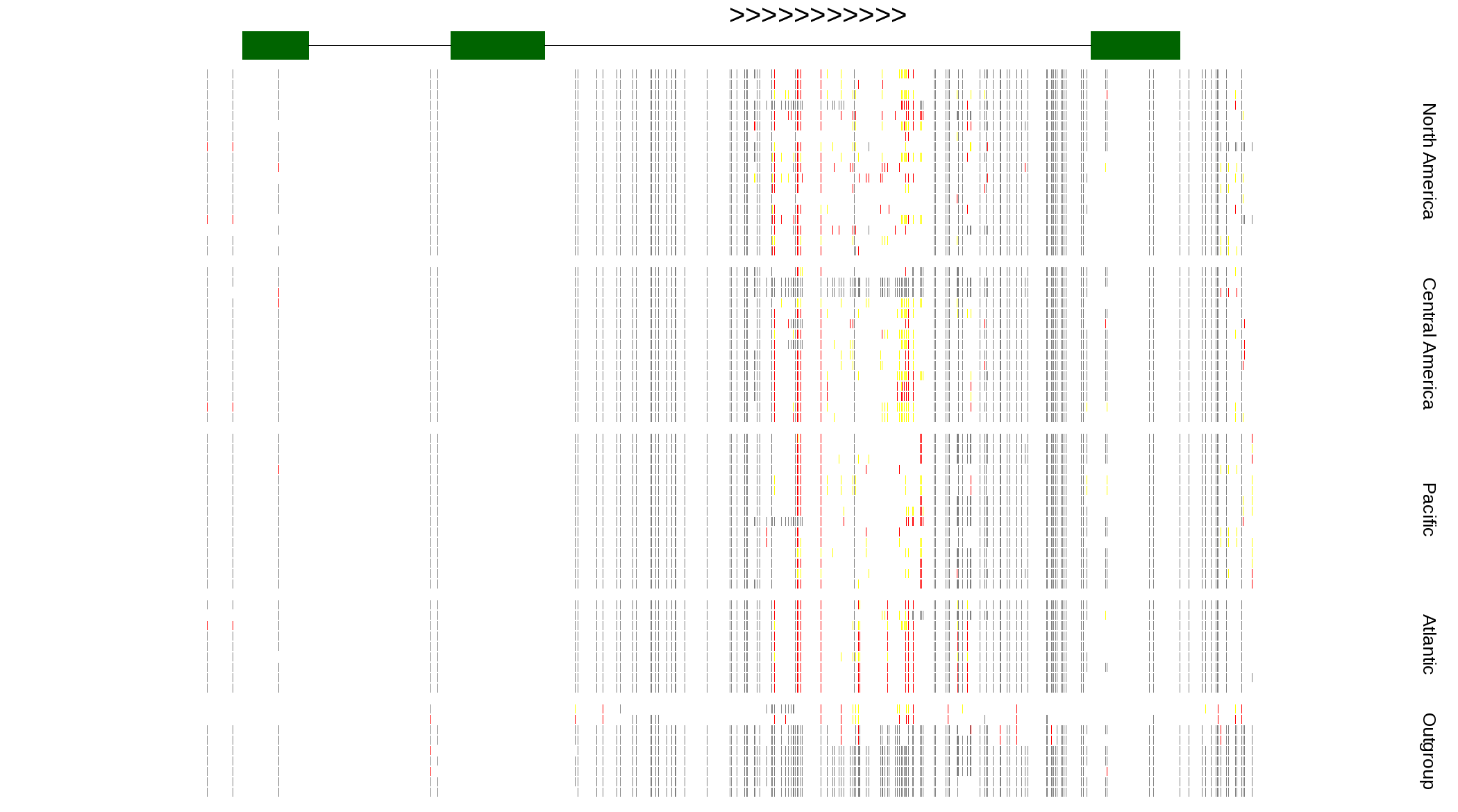
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS201570-TA
ATGTCAACGTCTACTATTTTGCGTGTAAAACGCCGGCTTGATGAAAACCCGCAAGATGCTTTAGTACTTGTTTGTAAGCGAATGAAAACGGATAAGGACGAAATATCCCCTTCACTATTTGTATTTAGAGGAAGTGTTGAAAACCAGAAGATCCGTAAGGAAAGAAAAGATGTTGCGAATGAGAGTCGTTATGAAATTGTGAATTGTAGCCGCGGCCTCGCTCAGTGCAACTCTGACAATTATGATCTAGTTGATCTGGAAAAGAAAGATGGGAAAGAAGACGCACAGTATACCTACGATCTGTATACACCGGTCAAGGAGGACTTTGATATAGCCATGTTGGATAATCTTGTCAGTATTGAGGATTACAGAACGGATCTCGTATTTGGTACGTATCGGGATTGTGATAATGTGGAGTCTGATGAAGACAGCGAAGATTCCAACGCTGAGAACGACTGGAGGAATGAATACCCTGACTCGGATCCCAGCAGCATCGATGAGGGGGACATGATAAAGGCTGTGGAGAACTGTAATATAGGTAAGAACTGCGAATAG
>DPOGS201570-PA
MSTSTILRVKRRLDENPQDALVLVCKRMKTDKDEISPSLFVFRGSVENQKIRKERKDVANESRYEIVNCSRGLAQCNSDNYDLVDLEKKDGKEDAQYTYDLYTPVKEDFDIAMLDNLVSIEDYRTDLVFGTYRDCDNVESDEDSEDSNAENDWRNEYPDSDPSSIDEGDMIKAVENCNIGKNCE-