| DPOGS202004 | ||
|---|---|---|
| Transcript | DPOGS202004-TA | 582 bp |
| Protein | DPOGS202004-PA | 193 aa |
| Genomic position | DPSCF300060 + 576776-578059 | |
| RNAseq coverage | 3916x (Rank: top 3%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006584 | 1e-91 | 92.81% | |
| Bombyx | BGIBMGA010409-TA | 8e-90 | 91.02% | |
| Drosophila | CG5023-PA | 4e-84 | 85.21% | |
| EBI UniRef50 | UniRef50_G6DP71 | 4e-94 | 100.00% | Calponin/transgelin n=10 Tax=Arthropoda RepID=G6DP71_DANPL |
| NCBI RefSeq | XP_001659123.1 | 7e-86 | 88.76% | calponin/transgelin [Aedes aegypti] |
| NCBI nr blastp | gi|157118472 | 1e-84 | 88.76% | calponin/transgelin [Aedes aegypti] |
| NCBI nr blastx | gi|157118472 | 4e-83 | 90.36% | calponin/transgelin [Aedes aegypti] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005515 | 7.1e-54 | protein binding | |
| GO:0003779 | 3.6e-10 | actin binding | ||
| GO:0031032 | 3.6e-10 | actomyosin structure organization | ||
| KEGG pathway | ptr:452604 | 2e-12 | ||
| K06084 (LMO7, FBXO20) | maps-> | Adherens junction | ||
| InterPro domain | [27-179] IPR001715 | 7.1e-54 | Calponin homology domain | |
| [28-41] IPR003096 | 3.5e-25 | Smooth muscle protein/calponin | ||
| [34-48] IPR001997 | 3.6e-10 | Calponin | ||
| [169-192] IPR000557 | 8.3e-08 | Calponin repeat | ||
| Orthology group | MCL15942 | Insect specific | ||
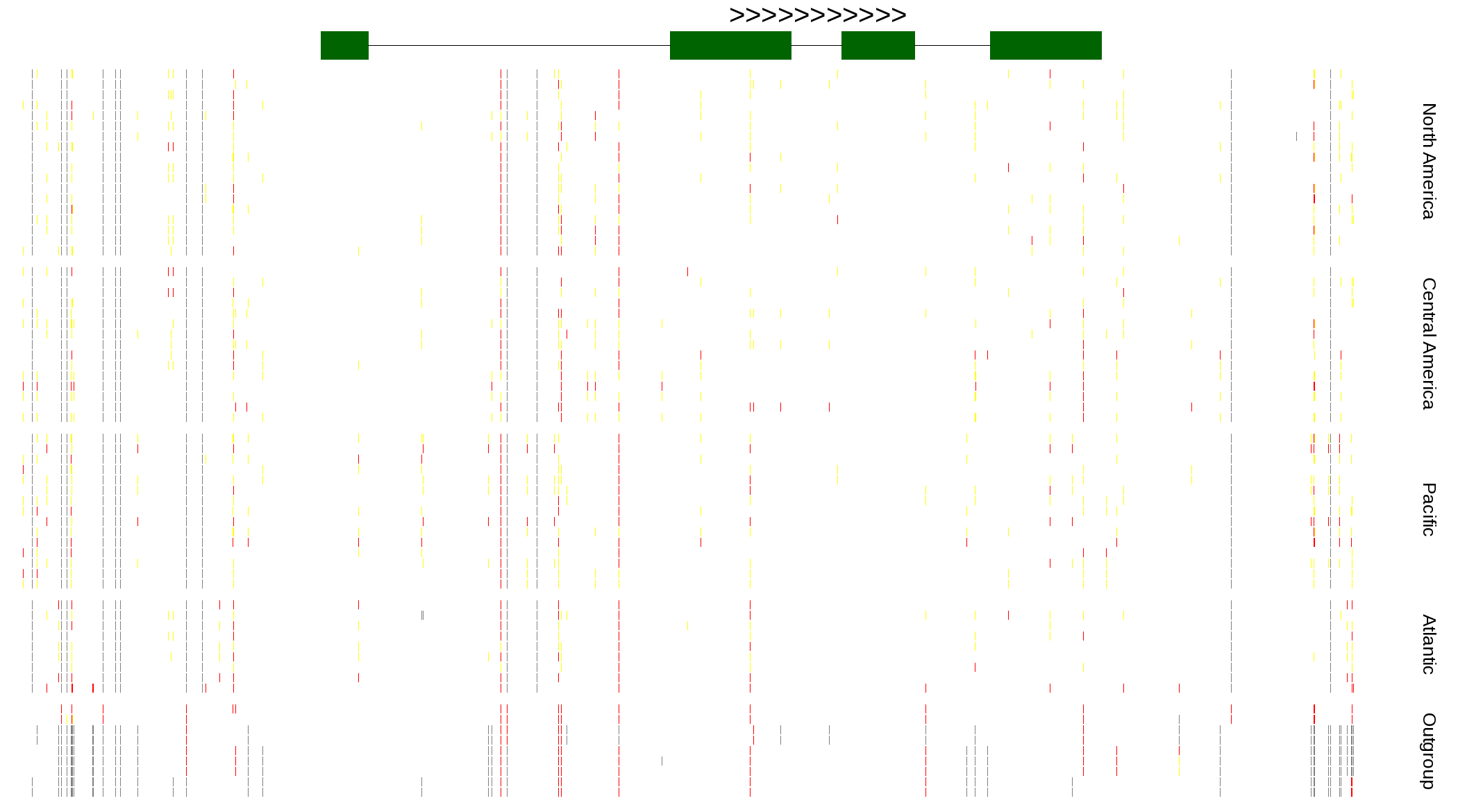
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS202004-TA
ATGGTACTTGTGTTGGAAGCTGTACCGACGCCAGCCGCCCCCGCGGACGCTGACCGCCTTTGGGCGGGCGGCTTTAGTGCCAGAAACAAGGAACAGGAGGAAGAAGTTCTAAACTGGATCAGTGCTGTTCTTGGGGAACCGCTTCCTAAAGGCGCGTATGAAGACATCTTGAAAGATGGAGTCATTCTGTGCAAGCTTATCAATAAATTGGCACCCGGATCAGTTAAAAAGATCCAGGAAAGGGGAACCAACTTTCAACTTATGGAAAACATTCAAAGGTTTCAAGCAGCTATTAAAAAATACGGAGTTCCAGAGGAGGAAATATTCCAGACAGCCGATCTTTTCGAGAGACGTAACATTCCTCAAGTCACCTTGTGCTTATACTCTCTCGGTAGAATAACACAGAAGCATCCCGAATATAACGGTCCTCAATTAGGACCTAAGATGGCGGAGAAGAATGAAAGATCTTTCACAGAAGAACAACTTCGGGCTCACAACGCTGAATTAAACCTACAAATGGGTTACAACAAGGGGGCGTCCCAATCGGGCCATGGTGGTTTTGGCAACACTCGTCATATGTAA
>DPOGS202004-PA
MVLVLEAVPTPAAPADADRLWAGGFSARNKEQEEEVLNWISAVLGEPLPKGAYEDILKDGVILCKLINKLAPGSVKKIQERGTNFQLMENIQRFQAAIKKYGVPEEEIFQTADLFERRNIPQVTLCLYSLGRITQKHPEYNGPQLGPKMAEKNERSFTEEQLRAHNAELNLQMGYNKGASQSGHGGFGNTRHM-