| DPOGS202301 | ||
|---|---|---|
| Transcript | DPOGS202301-TA | 492 bp |
| Protein | DPOGS202301-PA | 163 aa |
| Genomic position | DPSCF300032 + 219364-220244 | |
| RNAseq coverage | 774x (Rank: top 17%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL003447 | 2e-46 | 97.73% | |
| Bombyx | BGIBMGA004972-TA | 3e-57 | 97.12% | |
| Drosophila | pnt-PB | 3e-18 | 49.35% | |
| EBI UniRef50 | UniRef50_E2B8A7 | 3e-34 | 55.15% | ETS-like protein pointed, isoform P2 n=3 Tax=Formicidae RepID=E2B8A7_HARSA |
| NCBI RefSeq | XP_971341.2 | 1e-29 | 66.67% | PREDICTED: similar to Ets domain-containing protein [Tribolium castaneum] |
| NCBI nr blastp | gi|307212301 | 1e-33 | 55.15% | ETS-like protein pointed, isoform P2 [Harpegnathos saltator] |
| NCBI nr blastx | gi|307212301 | 2e-33 | 55.15% | ETS-like protein pointed, isoform P2 [Harpegnathos saltator] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005634 | 3.7e-25 | nucleus | |
| GO:0043565 | 3.7e-25 | sequence-specific DNA binding | ||
| GO:0005515 | 1.6e-22 | protein binding | ||
| KEGG pathway | tca:659984 | 3e-29 | ||
| K02678 (ETS, pnt) | maps-> | MAPK signaling pathway - fly | ||
| Dorso-ventral axis formation | ||||
| InterPro domain | [11-95] IPR003118 | 3.7e-25 | Sterile alpha motif/pointed | |
| [23-95] IPR013761 | 6.3e-24 | Sterile alpha motif-type | ||
| [5-98] IPR010993 | 1.6e-22 | Sterile alpha motif homology | ||
| Orthology group | MCL17778 | Insect specific | ||
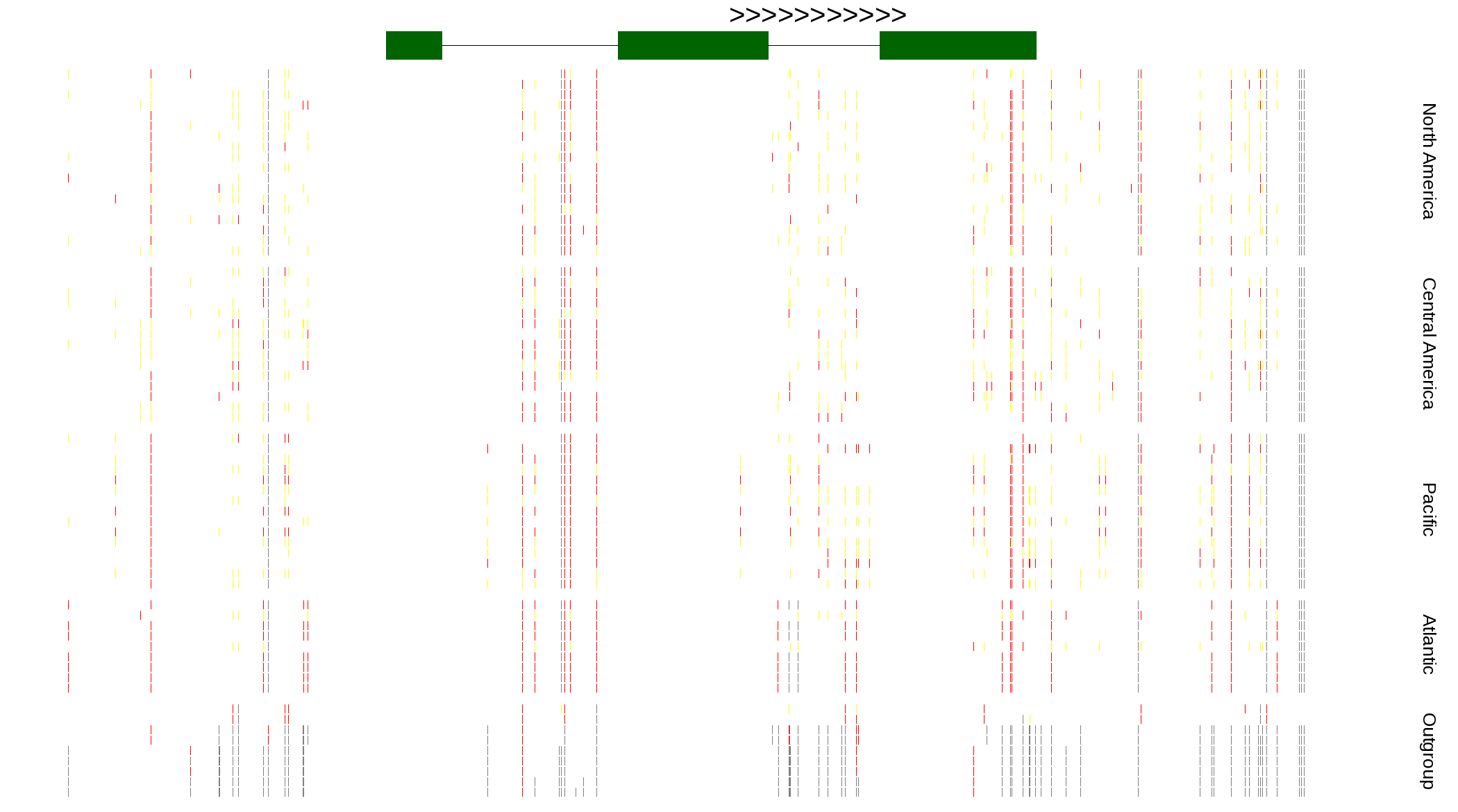
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS202301-TA
ATGTGGAAAGAATTAGAAATCACTTTTATCGTCCAGAGGCATGTAGTTCCAAGGTCGAGCTGCTTCCTGTGCGTGCACCCTCGTCAGTGGAGTGAGGCGGCTGTTGCTGCATGGCTCCGTTGGGCGGCTCGTGAGTTCTCCCTAGAAGGCGTAGCTCTACAGCAGTTCGCGCGTGCTCAAGGCAAGGATATATGCGCGATGGGAAGAGAGGAATTCGTGGCCAGAGCACCCGCTTTCATGGGCGATATACTTTGGGAACACCTGGAGATTCTACAAAAAGACGTGGAGAAGGAACGCTCTCTGCTGGCGAACGTGCCTCCCAATATGTATGAAAGCAACGTCTGTCTGCCGGAACTGGCCGACTACATCCCGCCGCCCGCCCACCACTACAACAACAATAACAATAACACAGGTATGCCCATTATAATAAAACACCAACCAGAGATGCTACCAGCTGATAGCTGTCAAAGATTGCAACGATACCATAAATAA
>DPOGS202301-PA
MWKELEITFIVQRHVVPRSSCFLCVHPRQWSEAAVAAWLRWAAREFSLEGVALQQFARAQGKDICAMGREEFVARAPAFMGDILWEHLEILQKDVEKERSLLANVPPNMYESNVCLPELADYIPPPAHHYNNNNNNTGMPIIIKHQPEMLPADSCQRLQRYHK-