| DPOGS202697 | ||
|---|---|---|
| Transcript | DPOGS202697-TA | 582 bp |
| Protein | DPOGS202697-PA | 193 aa |
| Genomic position | DPSCF300324 - 9289-10838 | |
| RNAseq coverage | 799x (Rank: top 16%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL021452 | 2e-48 | 45.79% | |
| Bombyx | BGIBMGA004865-TA | 4e-24 | 35.60% | |
| Drosophila | NLaz-PA | 1e-09 | 31.61% | |
| EBI UniRef50 | UniRef50_P09464 | 1e-34 | 45.45% | Bilin-binding protein n=5 Tax=Obtectomera RepID=BBP_PIEBR |
| NCBI RefSeq | NP_001036872.1 | 9e-37 | 44.50% | Bombyrin [Bombyx mori] |
| NCBI nr blastp | gi|112983654 | 1e-35 | 44.50% | Bombyrin precursor [Bombyx mori] |
| NCBI nr blastx | gi|270298182 | 1e-39 | 45.83% | bilin-binding protein [Pieris rapae] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005488 | 5e-45 | binding | |
| GO:0006810 | 1.2e-16 | transport | ||
| GO:0005215 | 1.2e-16 | transporter activity | ||
| KEGG pathway | ||||
| InterPro domain | [3-190] IPR011038 | 1.9e-46 | Calycin-like | |
| [1-193] IPR022271 | 4.4e-45 | Lipocalin, ApoD type | ||
| [9-191] IPR012674 | 5e-45 | Calycin | ||
| [40-180] IPR000566 | 1.4e-22 | Lipocalin/cytosolic fatty-acid binding protein domain | ||
| [31-47] IPR003057 | 1.2e-16 | Invertebrate colouration protein | ||
| Orthology group | MCL18582 | Lepidoptera specific | ||
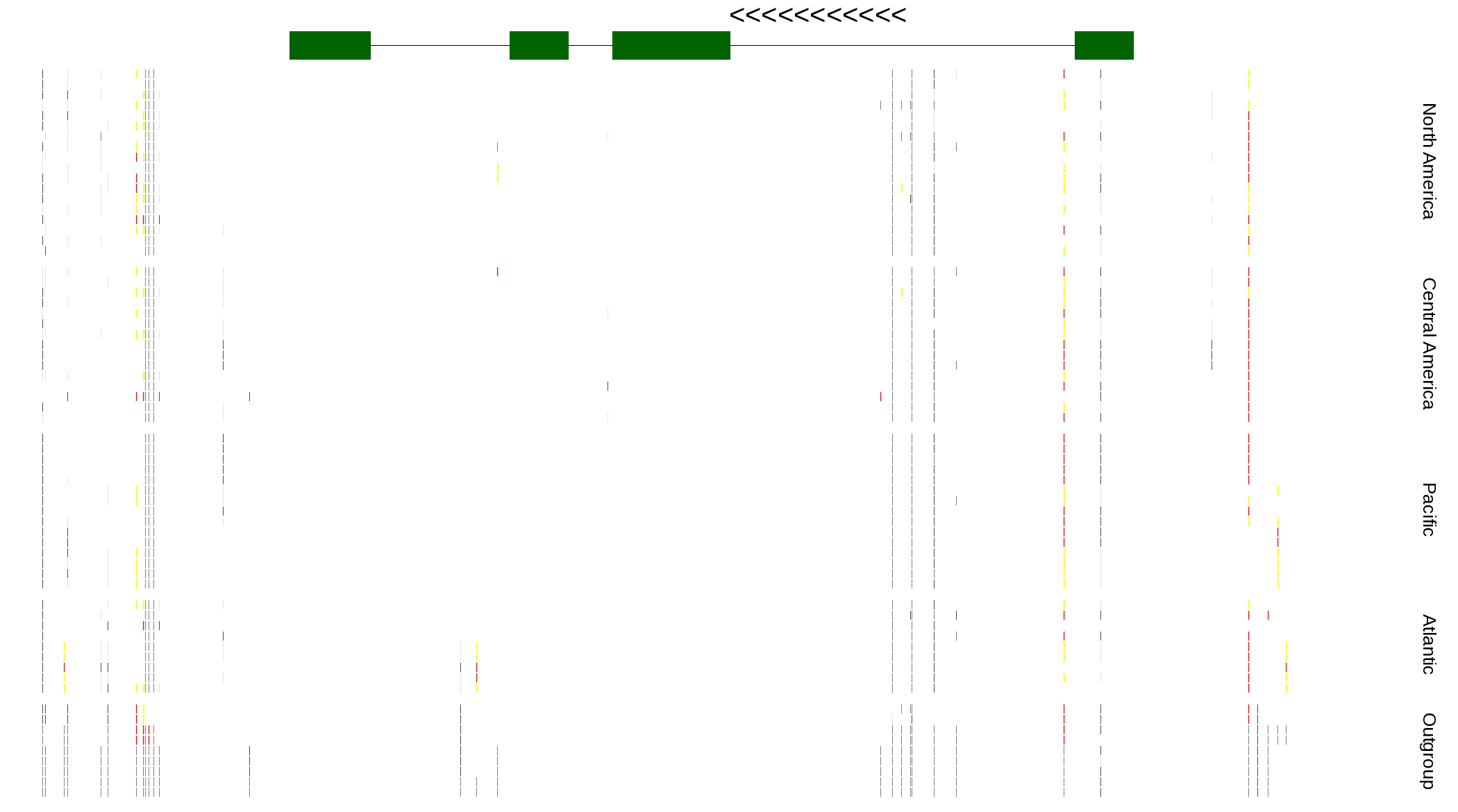
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS202697-TA
ATGTACGCTGTTGTTTTTCTAGCACTCGTCGCCTCAGCTCTTGCTGAAGTGTACGTTGATTCAAGATGCCCAGACATCCAGCCAGTTCAGAACTTCGACGTTGCTAATTACGGAAAGGGAGTCTGGTACGAAATAGCCAGGTACCCCAACAGGAAAGAAATGAAGATTGATTGCGGCCACACCGAGTATACTCCTCAAGGAGATCATTTTTCTGTAAAGAACTTGGGCTTTAATAACGGCAAACTTATGTCCATTGAGGGTGTGGCAAAACTTGCCGAGGACGCTGGAAATTCTGGAAAACTCATCTTTACCTTGCCATATGGAGCATCTGGCAAAAAAACTGACAACGTCTTGAATGTATTGTACACTGACTACGACAACTTCGCAATCGTCTATCACTGCCAGTTTCACGAGGAGAAGAACAGCCGTCAAGACTTTGCTTGGATTCTTTCGAGGTCTAAGTCCTTGTCCCCTGAATTGAAGGCTAAGGTCGACAAGTTCGTCTCGGAGTCCAAGGTTCTGGATAGCAGCAAATTCGTATGGCCCGACTTCTCTGACAAAGCCTGCAAGGCATCAGCTTAG
>DPOGS202697-PA
MYAVVFLALVASALAEVYVDSRCPDIQPVQNFDVANYGKGVWYEIARYPNRKEMKIDCGHTEYTPQGDHFSVKNLGFNNGKLMSIEGVAKLAEDAGNSGKLIFTLPYGASGKKTDNVLNVLYTDYDNFAIVYHCQFHEEKNSRQDFAWILSRSKSLSPELKAKVDKFVSESKVLDSSKFVWPDFSDKACKASA-