| DPOGS202895 | ||
|---|---|---|
| Transcript | DPOGS202895-TA | 636 bp |
| Protein | DPOGS202895-PA | 211 aa |
| Genomic position | DPSCF300126 - 210970-212779 | |
| RNAseq coverage | 78x (Rank: top 65%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL004898 | 2e-65 | 85.71% | |
| Bombyx | BGIBMGA004169-TA | 3e-83 | 80.00% | |
| Drosophila | CG30392-PA | 3e-48 | 46.92% | |
| EBI UniRef50 | UniRef50_F4WSH1 | 2e-49 | 45.41% | Glycolipid transfer protein domain-containing protein 1 n=6 Tax=Formicidae RepID=F4WSH1_ACREC |
| NCBI RefSeq | XP_316032.4 | 2e-56 | 51.47% | AGAP005990-PA [Anopheles gambiae str. PEST] |
| NCBI nr blastp | gi|158295135 | 3e-55 | 51.47% | AGAP005990-PA [Anopheles gambiae str. PEST] |
| NCBI nr blastx | gi|158295135 | 1e-52 | 51.47% | AGAP005990-PA [Anopheles gambiae str. PEST] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0017089 | 4.5e-53 | glycolipid transporter activity | |
| GO:0051861 | 4.5e-53 | glycolipid binding | ||
| GO:0046836 | 4.5e-53 | glycolipid transport | ||
| GO:0005737 | 4.5e-53 | cytoplasm | ||
| KEGG pathway | ||||
| InterPro domain | [16-211] IPR014830 | 4.5e-53 | Glycolipid transfer protein domain | |
| Orthology group | MCL11630 | Multiple-copy universal gene | ||
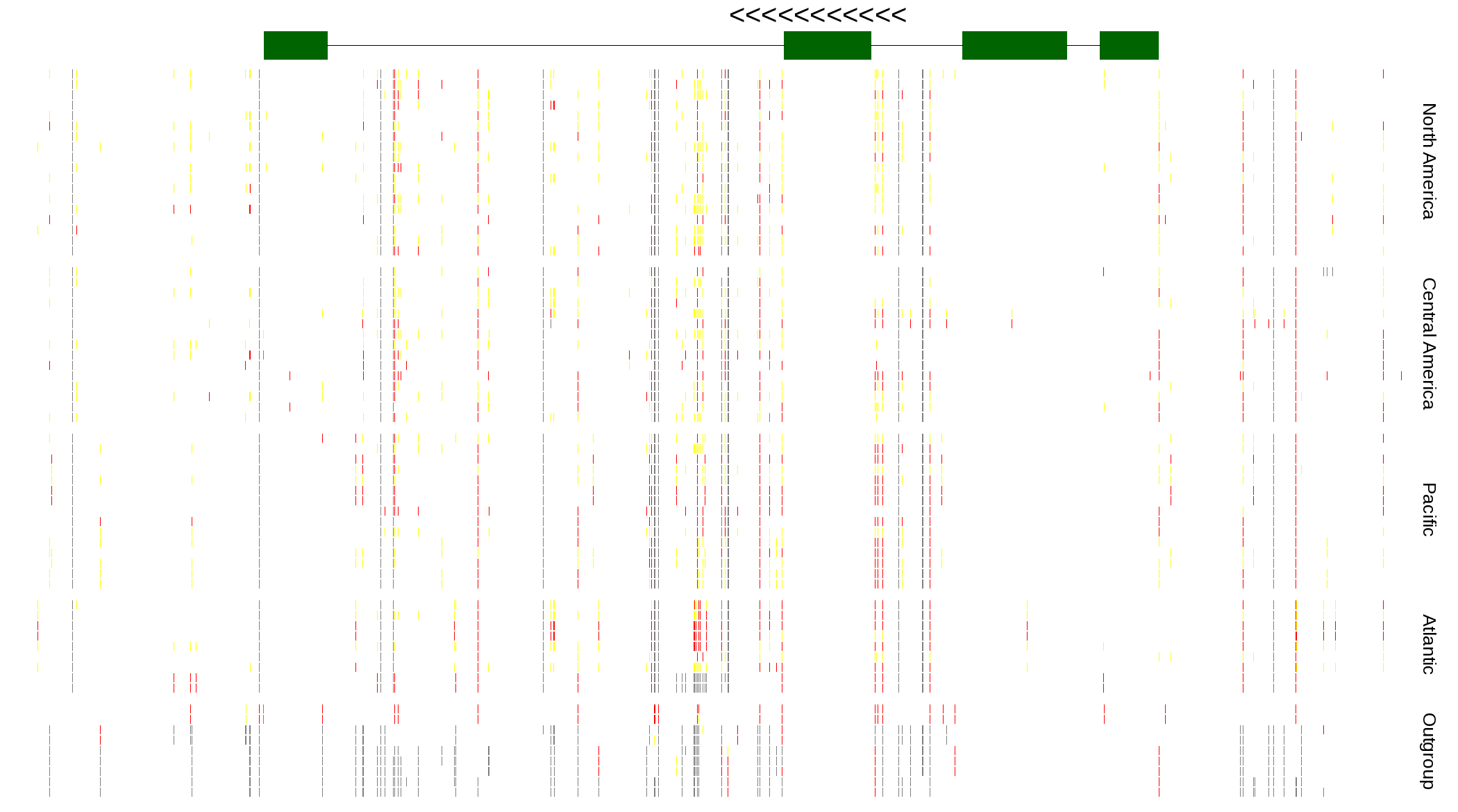
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS202895-TA
ATGTCCAATGAAAACACTTTAAATATACAATATGTACATGAAAGTTTTCAGAGAAGTTTAAAAGAAGACGACGATGTGATTATAGACGATTATATAGAAGGTTACAATGAACTAGTAAAGTTCCTGAATCTCATCGGTTCAGTTTTTTCTTTTGTAAGCAGCGATGTGCGAAGTAAAATAAAAATTATGGAGAAACATCGCGAAGGCGATGATTCTCTTTACTTTGAATCCTTTAAGAAAATGATGAAATATGAAAAGGAAACAAGCTTACACGAAAAGAATGGCTACGTTTCTGGATCGAGGACAATGCTGAGATTACACAGAGGCTTAGATTTTATACGTCTGTTTCTTAAGAGACTGAGTGAAGCTGAAGAGAGTATGAACACATGTACAACATGTCAGAACTCATACAATGAAACACTTGCTGCATTCCATCCCTGGTACATACGGAAGGCGGCCACTTTAGCTATGCATGCTTTGCCTTCCCGACCTGATTTATTAAAGAAGATATTCGGAACGGAAGAACAACTTGCGGAGGCTTTAAATATTTTACCACAGACTCTACAATCATGTGATGAGGTTTACAACCGGGTCGAGAAATTGTACACAGATTTCGATTTCCACGGCCTGCCGTAA
>DPOGS202895-PA
MSNENTLNIQYVHESFQRSLKEDDDVIIDDYIEGYNELVKFLNLIGSVFSFVSSDVRSKIKIMEKHREGDDSLYFESFKKMMKYEKETSLHEKNGYVSGSRTMLRLHRGLDFIRLFLKRLSEAEESMNTCTTCQNSYNETLAAFHPWYIRKAATLAMHALPSRPDLLKKIFGTEEQLAEALNILPQTLQSCDEVYNRVEKLYTDFDFHGLP-