| DPOGS203006 | ||
|---|---|---|
| Transcript | DPOGS203006-TA | 462 bp |
| Protein | DPOGS203006-PA | 153 aa |
| Genomic position | DPSCF300068 + 86658-89361 | |
| RNAseq coverage | 1030x (Rank: top 12%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL011018 | 8e-64 | 71.43% | |
| Bombyx | BGIBMGA003865-TA | 6e-53 | 59.75% | |
| Drosophila | ppl-PA | 2e-38 | 56.74% | |
| EBI UniRef50 | UniRef50_E2BX40 | 4e-43 | 65.32% | Glycine cleavage system H protein, mitochondrial n=19 Tax=cellular organisms RepID=E2BX40_HARSA |
| NCBI RefSeq | XP_001866806.1 | 1e-44 | 56.49% | glycine cleavage system h protein [Culex quinquefasciatus] |
| NCBI nr blastp | gi|307169516 | 7e-44 | 65.57% | Glycine cleavage system H protein, mitochondrial [Camponotus floridanus] |
| NCBI nr blastx | gi|307169516 | 2e-42 | 65.57% | Glycine cleavage system H protein, mitochondrial [Camponotus floridanus] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005960 | 1e-73 | glycine cleavage complex | |
| GO:0006546 | 1e-73 | glycine catabolic process | ||
| GO:0019464 | 1.5e-44 | glycine decarboxylation via glycine cleavage system | ||
| KEGG pathway | ||||
| InterPro domain | [30-154] IPR002930 | 1e-73 | Glycine cleavage H-protein | |
| [34-151] IPR017453 | 1.5e-44 | Glycine cleavage H-protein, subgroup | ||
| [27-153] IPR011053 | 6.3e-42 | Single hybrid motif | ||
| Orthology group | MCL11460 | Multiple-copy universal gene | ||
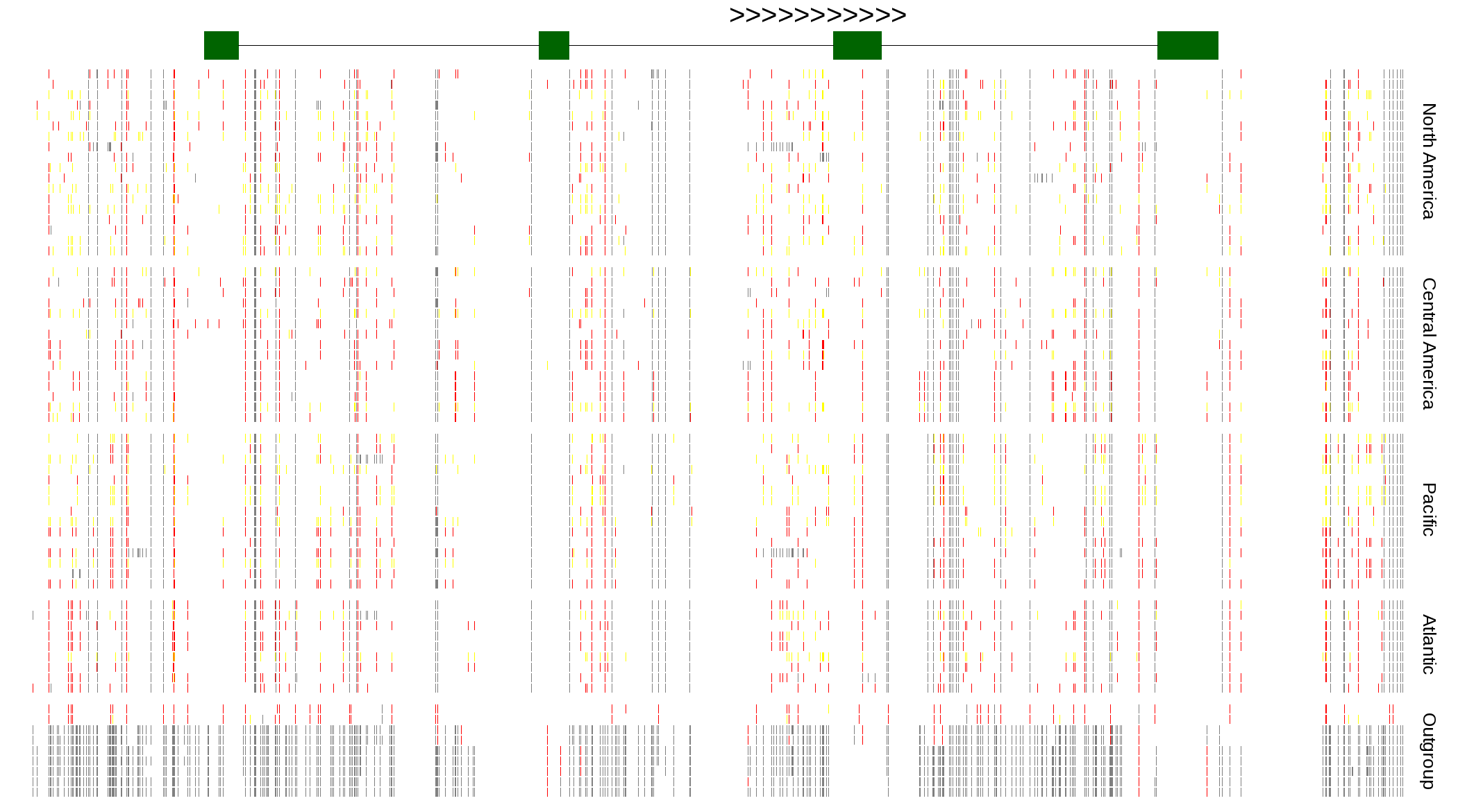
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS203006-TA
ATGGTGTTAGGAAAATTACTACGGTCCACGGCGCGTTTTGCTGGAGCGACACAATGCTGTGGCCGGGCCGTTTACAGTACTCAAGTTAAAGGAAGACTCTATACAAAGAAACACGAGTGGGTGTCTTTGGACGGCGGCGTAGGCACCGTTGGCATTTCTAATTATGCACAGGATGCTCTCGGGGAGGTTGTATTTGTGCAGTTGCCGGAGGTAGGTCGGACAGTAGCTGCTGCTGAGGAAGTCGGTGCTTTGGAGAGTGTCAAAGCTGCCAGTGAGGTCTACTCTCCTATATCAGGGACTATAACTGAGAAGAATGACGCGGTGGAGAGCAGTCCGTCCCTCATCAACAAGTCTTGTTATGAAGATGGCTGGCTGTTCAAGGTCTCGCTGAGCAAGCCAGATGAACTGGAGTCGCTCATGGACCAGCAGACCTACGACAAGTACCTCGAGGATTCACACTGA
>DPOGS203006-PA
MVLGKLLRSTARFAGATQCCGRAVYSTQVKGRLYTKKHEWVSLDGGVGTVGISNYAQDALGEVVFVQLPEVGRTVAAAEEVGALESVKAASEVYSPISGTITEKNDAVESSPSLINKSCYEDGWLFKVSLSKPDELESLMDQQTYDKYLEDSH-