| DPOGS203181 | ||
|---|---|---|
| Transcript | DPOGS203181-TA | 648 bp |
| Protein | DPOGS203181-PA | 215 aa |
| Genomic position | DPSCF300035 - 170652-173225 | |
| RNAseq coverage | 13x (Rank: top 82%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL021548 | 2e-55 | 45.97% | |
| Bombyx | BGIBMGA011172-TA | 2e-79 | 63.03% | |
| Drosophila | CG3829-PA | 5e-36 | 35.47% | |
| EBI UniRef50 | UniRef50_D2A3T6 | 5e-46 | 38.53% | Putative uncharacterized protein GLEAN_14951 n=1 Tax=Tribolium castaneum RepID=D2A3T6_TRICA |
| NCBI RefSeq | NP_001164133.1 | 9e-47 | 38.53% | scavenger receptor protein [Tribolium castaneum] |
| NCBI nr blastp | gi|282165788 | 2e-45 | 38.53% | scavenger receptor protein [Tribolium castaneum] |
| NCBI nr blastx | gi|282165788 | 2e-44 | 38.53% | scavenger receptor protein [Tribolium castaneum] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016020 | 2.8e-54 | membrane | |
| GO:0007155 | 2.8e-54 | cell adhesion | ||
| KEGG pathway | dre:192340 | 8e-25 | ||
| K12384 (SCARB2, LIMP2, CD36L2) | maps-> | Lysosome | ||
| InterPro domain | [1-198] IPR002159 | 2.8e-54 | CD36 antigen | |
| Orthology group | MCL10509 | Multiple-copy universal gene | ||
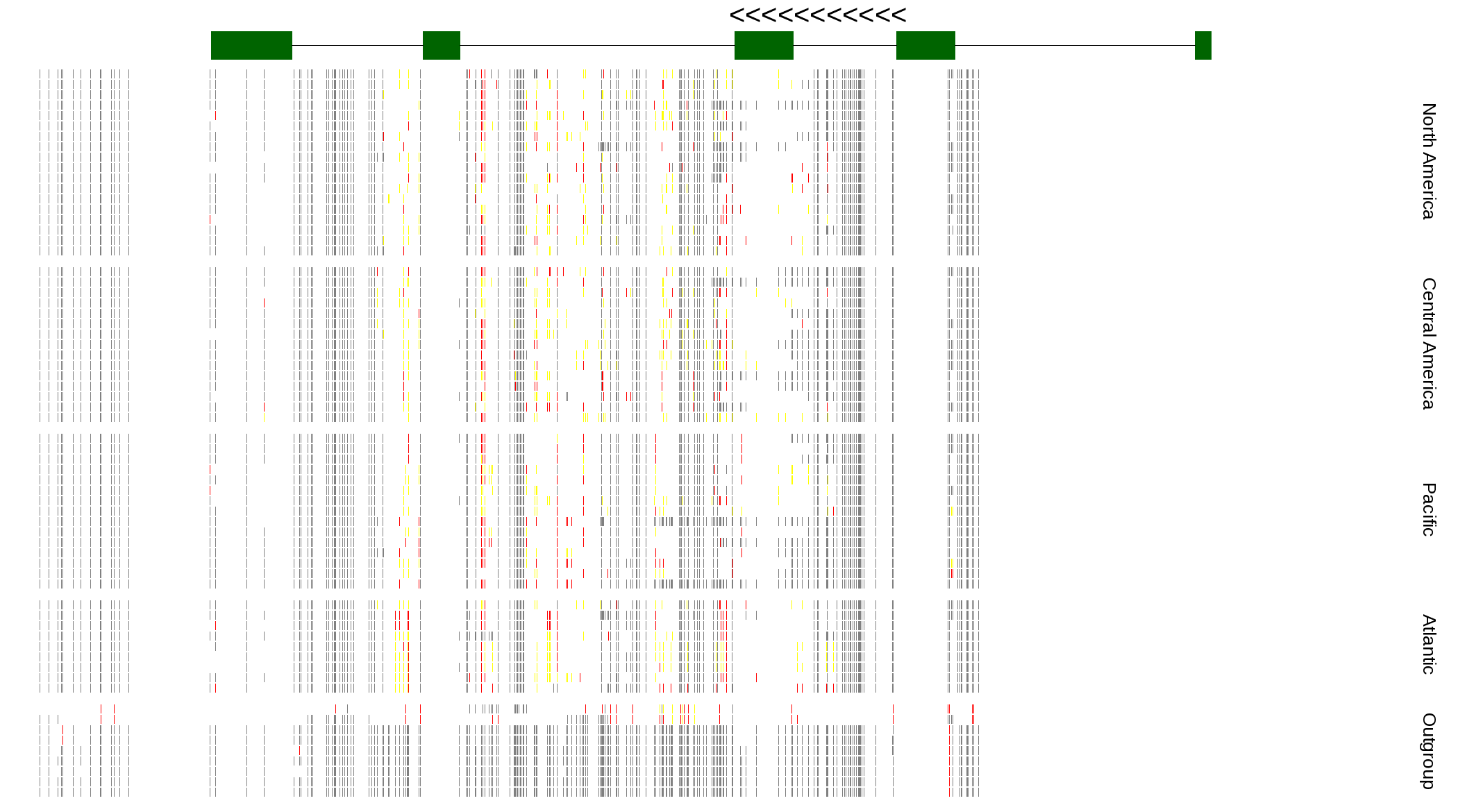
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS203181-TA
ATGTGCAGGAGACTGCCGTTCGACTACCTCAAGGACGTTGAGATGGAACAAGGGATTCGTCTCATGAGGTATGTCATGCCATCAAACGTGTTCGACGATCCCCAAAGCAATCCCGACAACCAGTGCTACTGTGATGTGGACAGCGGTACCTGCCCTCCGAGAGGAATCATAAACGTCACAGCCTGTTCTATGGGCGCTCCCCTGGTGGCTTCCTTTCCCCATTTCTACCGCGGAGACCCCAAACTGTATGAGGACATCCAAGGCCTGAGCCCCAACGGCGAGCTCCATGACTCCTTTATTGACATACACCCAACCCTTGGCATCGCACTGAACGGCAGATCAAGCCTCCAGCTGAACATCCAAGTCAAGAAGTCCAGCGTGTTCGGCGCTCTCAGTTTCCTGCCAGAAGGAATTATCCTGCCAATCGCGTGGATAGAAATGGCGCTCGAAGAACTACCAGAGAGTCTTCAATCGTTGGTCTACCACGGGACTTTTTCCACAGCGGCCGTGCAACTCGGCCTGGCCATGTTCTGTTCCGTAACCCTGCTCATTTCTTCCATATGCATGCTGATGATGATCATCACTAGAAGAAGGAAACCATGCGCCACCCTCAAAATTATACCAGCCGACATTGAACTGAAGACTTAA
>DPOGS203181-PA
MCRRLPFDYLKDVEMEQGIRLMRYVMPSNVFDDPQSNPDNQCYCDVDSGTCPPRGIINVTACSMGAPLVASFPHFYRGDPKLYEDIQGLSPNGELHDSFIDIHPTLGIALNGRSSLQLNIQVKKSSVFGALSFLPEGIILPIAWIEMALEELPESLQSLVYHGTFSTAAVQLGLAMFCSVTLLISSICMLMMIITRRRKPCATLKIIPADIELKT-