| DPOGS203259 | ||
|---|---|---|
| Transcript | DPOGS203259-TA | 687 bp |
| Protein | DPOGS203259-PA | 228 aa |
| Genomic position | DPSCF300229 - 198527-199213 | |
| RNAseq coverage | 24x (Rank: top 77%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006693 | 2e-75 | 57.33% | |
| Bombyx | BGIBMGA002640-TA | 2e-73 | 55.56% | |
| Drosophila | Uch-PA | 8e-62 | 46.93% | |
| EBI UniRef50 | UniRef50_E5EVW1 | 3e-71 | 55.56% | Ubiquitin carboxy-terminal hydrolase CG4265 n=11 Tax=Neoptera RepID=E5EVW1_BOMMO |
| NCBI RefSeq | NP_476940.1 | 2e-60 | 46.93% | ubiquitin carboxy-terminal hydrolase [Drosophila melanogaster] |
| NCBI nr blastp | gi|312597590 | 9e-71 | 55.56% | ubiquitin carboxy-terminal hydrolase CG4265 [Bombyx mori] |
| NCBI nr blastx | gi|312597590 | 2e-69 | 56.05% | ubiquitin carboxy-terminal hydrolase CG4265 [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0006511 | 6.2e-88 | ubiquitin-dependent protein catabolic process | |
| GO:0005622 | 6.2e-88 | intracellular | ||
| GO:0004221 | 6.2e-88 | ubiquitin thiolesterase activity | ||
| KEGG pathway | ||||
| InterPro domain | [1-228] IPR001578 | 6.2e-88 | Peptidase C12, ubiquitin carboxyl-terminal hydrolase 1 | |
| Orthology group | ||||
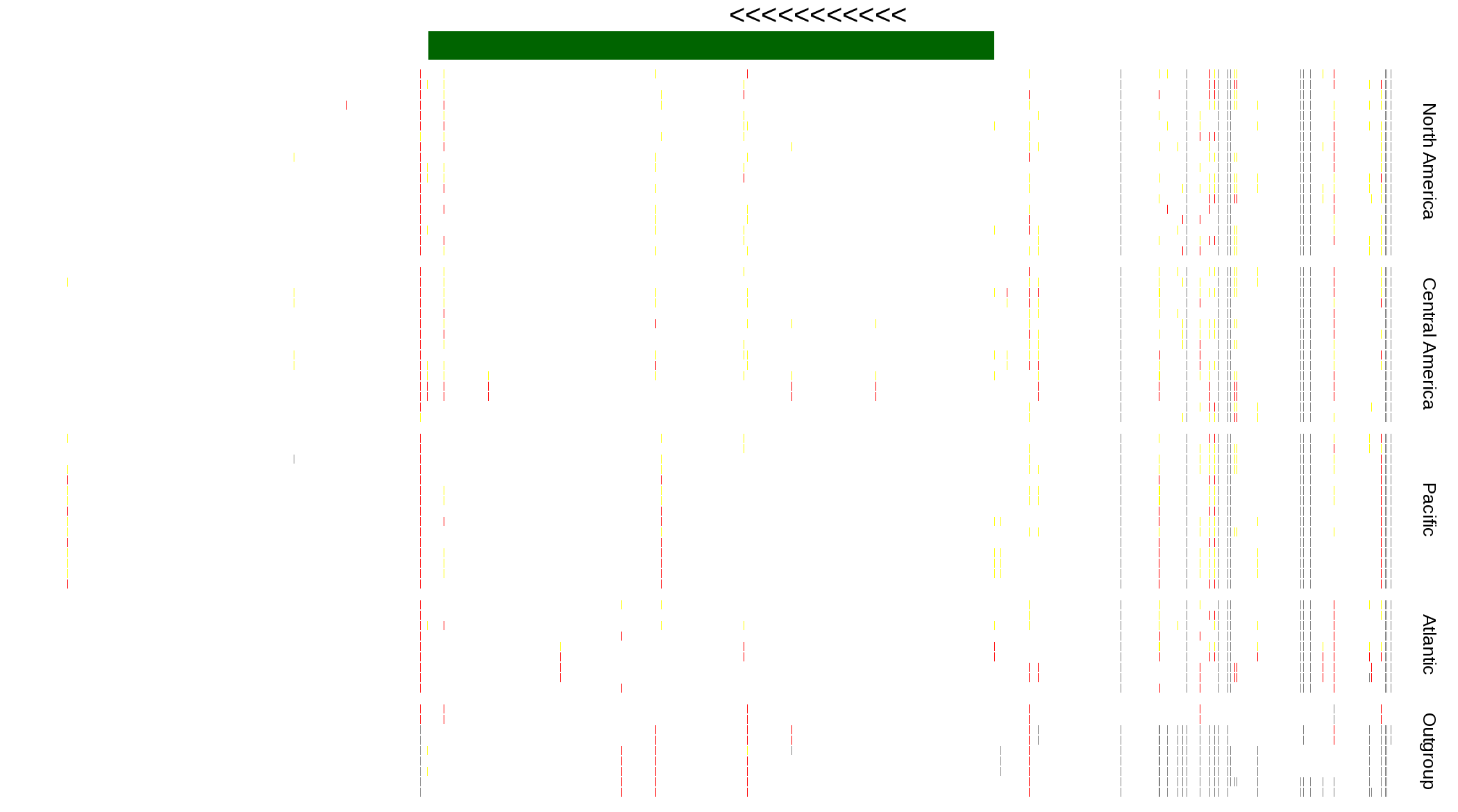
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS203259-TA
ATGGCTGTTTCTTTGCCTATGGAAGCAAATCCTGAAACTATGAACAAATACCTTCAGAGGCTTGGCGTTTCTAACAAATGGCGTATGGTTGATGTTATGGGTTTAGAAGGTGATCCTTTAAAATGGGTGCCACGGCCAGTTCTCGCTGTTATACTTTTATTTCCGCTTTCTGAGCATTATGAAAGACATCGCTCTCAGCAGGAGAACGAGCTACAAATCAAACCTCAGCAACCTCCTAAGGACGTGTTCCATTTAAAACAAGTGCTCAGTAATGTGTGTGGGACAATTGCTCTAGTGCACAGTGTCGCTAATAATATTAATACCATCGATTTAGAAGATGGTATTTTAAAAGATTACTTGAAGTCCGCTTCAGGCTTGGACGCAGCAGCTAAAGGCGCTCTTTTGGAGAATTCAACACATATTTTTAGGGCATATGAGGAAGTTTTAAACTCAGCGAATAATCCTGCAAATGAAGACAGCGAACCTGTGAACAATCATTTTATAACATTTATCCATAAGAATGGTAATCTTTATGAGTTGGATGGTCGTAAGTCAATACCTATTAACCACGGCCCAACGACTCCGGAAACTCTGTTGGATGACTCTGCCAAAATTTGCCTGGAATTCATAGCTCGCGAGCCAGATAATATAGGTTTTAATGTCGTTGCTCTCGTTCCTGCTCAATAA
>DPOGS203259-PA
MAVSLPMEANPETMNKYLQRLGVSNKWRMVDVMGLEGDPLKWVPRPVLAVILLFPLSEHYERHRSQQENELQIKPQQPPKDVFHLKQVLSNVCGTIALVHSVANNINTIDLEDGILKDYLKSASGLDAAAKGALLENSTHIFRAYEEVLNSANNPANEDSEPVNNHFITFIHKNGNLYELDGRKSIPINHGPTTPETLLDDSAKICLEFIAREPDNIGFNVVALVPAQ-