| DPOGS203654 | ||
|---|---|---|
| Transcript | DPOGS203654-TA | 606 bp |
| Protein | DPOGS203654-PA | 201 aa |
| Genomic position | DPSCF300010 - 2786057-2790315 | |
| RNAseq coverage | 592x (Rank: top 21%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006932 | 1e-96 | 84.10% | |
| Bombyx | BGIBMGA003451-TA | 2e-93 | 80.00% | |
| Drosophila | CG11811-PA | 2e-66 | 59.49% | |
| EBI UniRef50 | UniRef50_G6D291 | 5e-115 | 100.00% | Guanylate kinase n=4 Tax=Opisthokonta RepID=G6D291_DANPL |
| NCBI RefSeq | NP_001040302.1 | 6e-93 | 81.03% | guanylate kinase [Bombyx mori] |
| NCBI nr blastp | gi|114051493 | 9e-92 | 81.03% | guanylate kinase [Bombyx mori] |
| NCBI nr blastx | gi|114051493 | 3e-89 | 81.03% | guanylate kinase [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005515 | 2.5e-79 | protein binding | |
| GO:0006163 | 2.3e-60 | purine nucleotide metabolic process | ||
| GO:0004385 | 2.3e-60 | guanylate kinase activity | ||
| KEGG pathway | aga:AgaP_AGAP011105 | 2e-74 | ||
| K00942 (E2.7.4.8, gmk) | maps-> | Purine metabolism | ||
| InterPro domain | [5-191] IPR008145 | 2.5e-79 | Guanylate kinase/L-type calcium channel | |
| [7-189] IPR008144 | 5.9e-66 | Guanylate kinase | ||
| [8-188] IPR017665 | 2.3e-60 | Guanylate kinase, sub-group | ||
| Orthology group | MCL13408 | Single-copy universal gene | ||
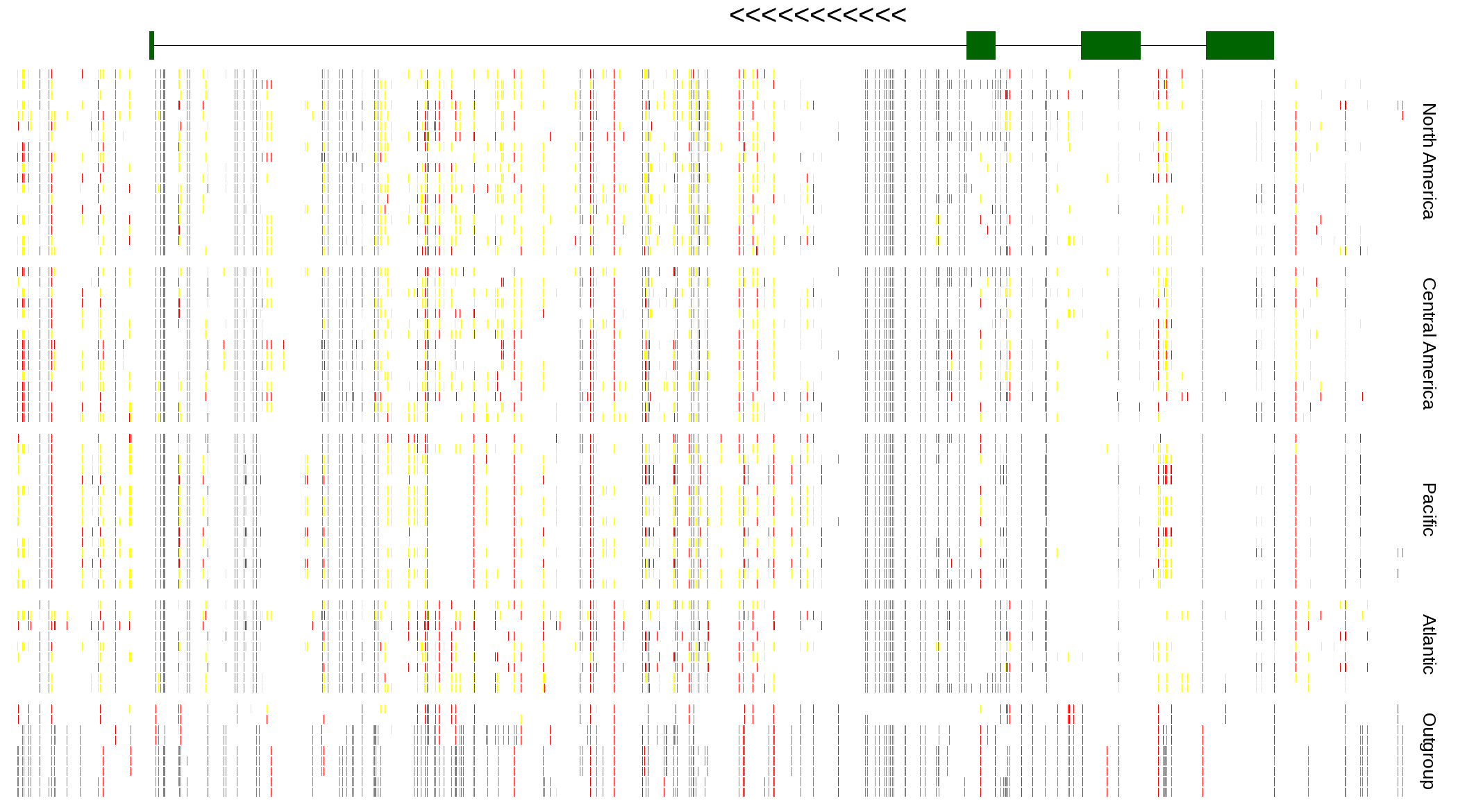
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS203654-TA
ATGGTGCAGAAAGGTCCTCGTCCCTTGGTTCTTTGTGGACCCTCCGGCTCAGGGAAAAGTACATTGCTGAAGCGGCTTTTAAACGATTTTCCTGACAAGTTCGGATTTGGTGTATCTCATACCACAAGAGGCCCTCGTCCTGGCGAAAAAGATGGTGTTCATTATCACTTTACTAATAAAAGTGACATGATTGCAGCAATTGAAAAAGGCGAATTTATAGAAACTGCAACATTTAGCGGAAATATGTATGGGACAAGCAAGAGAGCAGTAGAAGATGTCCGATGCACTGGCAAGACATGCATTTTAGACATAGAAATAGAGGGTGTAAAGCAGATTAAGAAGACGGATTTAGACCCACTACTGGTGTTTGTGATGCCACCATCCATTGAACAGCTTGAGAAACGTCTACGAGCTCGTAACACAGAACAGGAGGATGCATTAAAGAAAAGATTGGAAACGGCAAGTCGTGAGATTTTATATGGACAAGAACCCGGTAATTTCCATATAATCATTGTCAATGATAACTTAGACAAGGCGTATTCTGAGTTACATGATTTTGTGTCCCAAAACTTACAAAGTAGTGAACCAGCTCTTCGTGGACTTTAA
>DPOGS203654-PA
MVQKGPRPLVLCGPSGSGKSTLLKRLLNDFPDKFGFGVSHTTRGPRPGEKDGVHYHFTNKSDMIAAIEKGEFIETATFSGNMYGTSKRAVEDVRCTGKTCILDIEIEGVKQIKKTDLDPLLVFVMPPSIEQLEKRLRARNTEQEDALKKRLETASREILYGQEPGNFHIIIVNDNLDKAYSELHDFVSQNLQSSEPALRGL-