| DPOGS204008 | ||
|---|---|---|
| Transcript | DPOGS204008-TA | 498 bp |
| Protein | DPOGS204008-PA | 165 aa |
| Genomic position | DPSCF300470 + 44150-46017 | |
| RNAseq coverage | 350x (Rank: top 33%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL011247 | 3e-84 | 97.58% | |
| Bombyx | BGIBMGA006476-TA | 7e-10 | 41.10% | |
| Drosophila | tsu-PA | 1e-59 | 66.27% | |
| EBI UniRef50 | UniRef50_C1BGS0 | 9e-53 | 64.60% | RNA-binding protein 8A n=10 Tax=Metazoa RepID=C1BGS0_ONCMY |
| NCBI RefSeq | NP_001037578.1 | 2e-86 | 93.98% | RNA binding motif protein Y14 [Bombyx mori] |
| NCBI nr blastp | gi|301072399 | 1e-85 | 95.18% | RNA-binding protein [Helicoverpa armigera] |
| NCBI nr blastx | gi|301072399 | 1e-85 | 95.18% | RNA-binding protein [Helicoverpa armigera] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0000166 | 2.7e-22 | nucleotide binding | |
| GO:0003676 | 3e-20 | nucleic acid binding | ||
| GO:0006396 | 7.2e-20 | RNA processing | ||
| GO:0005634 | 7.2e-20 | nucleus | ||
| GO:0003723 | 7.2e-20 | RNA binding | ||
| GO:0005737 | 7.2e-20 | cytoplasm | ||
| KEGG pathway | nvi:100115053 | 1e-61 | ||
| K12876 (RBM8A, Y14) | maps-> | Spliceosome | ||
| InterPro domain | [67-157] IPR012677 | 2.7e-22 | Nucleotide-binding, alpha-beta plait | |
| [77-150] IPR000504 | 3e-20 | RNA recognition motif domain | ||
| [29-41] IPR008111 | 7.2e-20 | RNA-binding motif protein 8 | ||
| Orthology group | MCL14259 | Single-copy universal gene | ||
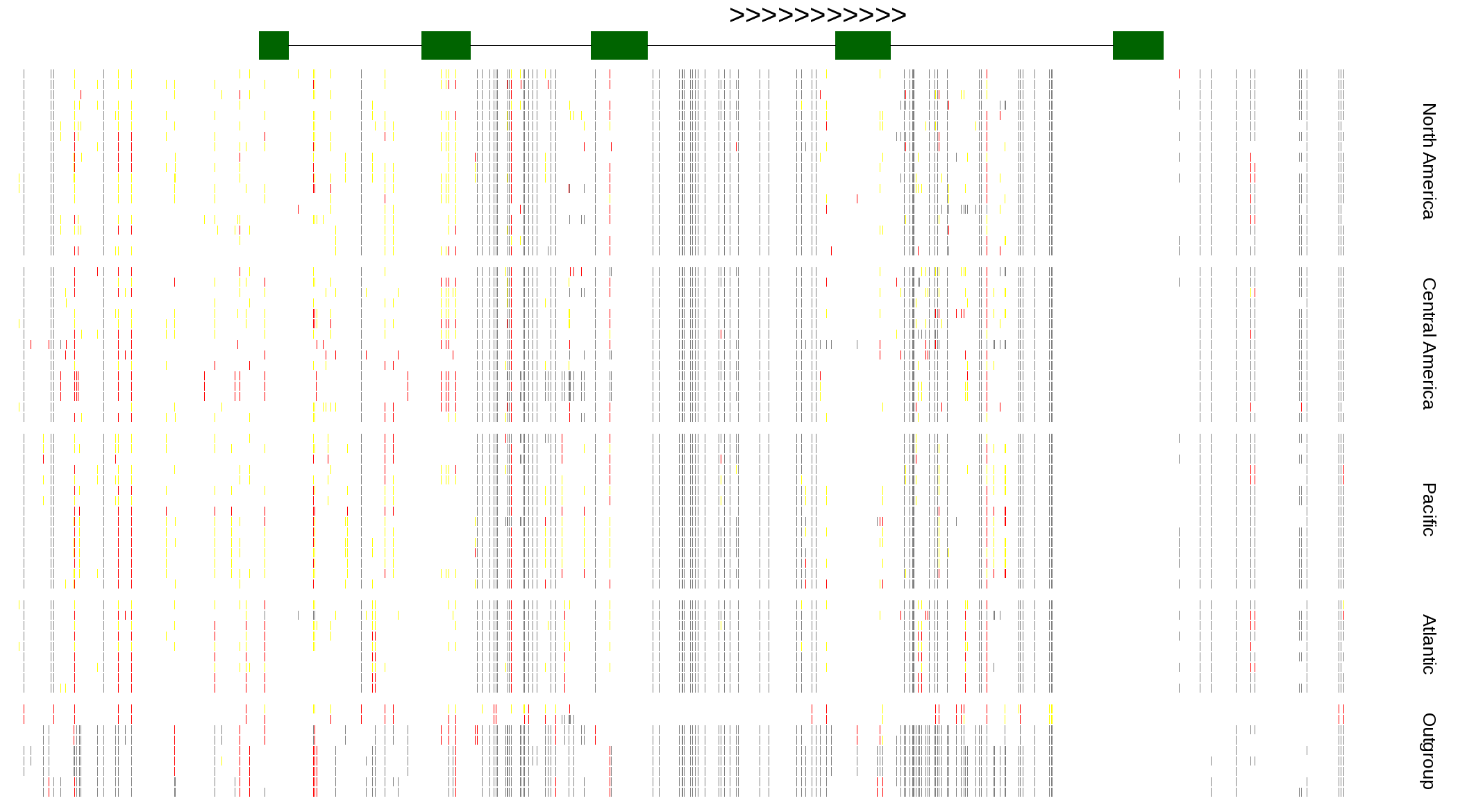
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204008-TA
ATGGCTGACGTGTTAGATATTGAAAATTCAGAAGAATTTGAAGTGGACGAAGATGGTGAACAGGGTATTGTCCGTCTGAAGGAAAAGGCGCGCAAACGTAAAGGTCGTGGATTTGGATCGGAGGCAGGCGGAGGCGGCTCCGAAAGAGGCAATAGGGGCAGATATGACAGCCTTGCACCAGAAGGTAATGGGAATACTCCTGGGCCACAAAGATCGGTTGAAGGATGGATCCTGTTTGTCAGTAATGTGCATGAAGAAGCACAGGAAGAAGATATTCAGAACCAGTTCTCAGAGTTTGGGGAGATAAAGAATATTCATTTAAATCTTGATAGACGCACGGGATTCTTGAAAGGCTATGCCCTAGTCGAGTACGAGACGTACAAACAGGCTGCTAGTGCTAGGGAAGCTCTAGACGGTACAGACATACTCGGTCAGGCAATCTCTGTGGACTGGTGTTTTGTTAAAGGACCAACCAAGACTCACAAGAAACGGCGATAA
>DPOGS204008-PA
MADVLDIENSEEFEVDEDGEQGIVRLKEKARKRKGRGFGSEAGGGGSERGNRGRYDSLAPEGNGNTPGPQRSVEGWILFVSNVHEEAQEEDIQNQFSEFGEIKNIHLNLDRRTGFLKGYALVEYETYKQAASAREALDGTDILGQAISVDWCFVKGPTKTHKKRR-