| DPOGS204021 | ||
|---|---|---|
| Transcript | DPOGS204021-TA | 150 bp |
| Protein | DPOGS204021-PA | 49 aa |
| Genomic position | DPSCF300138 - 162461-162610 | |
| RNAseq coverage | 10x (Rank: top 84%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL010644 | 6e-15 | 75.00% | |
| Bombyx | BGIBMGA004873-TA | 2e-19 | 83.67% | |
| Drosophila | CG11337-PD | 5e-14 | 75.00% | |
| EBI UniRef50 | UniRef50_A8JRH9 | 1e-11 | 75.00% | CG11337, isoform C n=16 Tax=Coelomata RepID=A8JRH9_DROME |
| NCBI RefSeq | XP_001651608.1 | 1e-14 | 78.72% | polyribonucleotide nucleotidyltransferase [Aedes aegypti] |
| NCBI nr blastp | gi|157111536 | 3e-13 | 78.72% | polyribonucleotide nucleotidyltransferase [Aedes aegypti] |
| NCBI nr blastx | gi|157111536 | 5e-13 | 79.17% | polyribonucleotide nucleotidyltransferase [Aedes aegypti] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003723 | 3.8e-13 | RNA binding | |
| GO:0004654 | 3.8e-13 | polyribonucleotide nucleotidyltransferase activity | ||
| GO:0006402 | 3.8e-13 | mRNA catabolic process | ||
| KEGG pathway | aag:AaeL_AAEL000922 | 3e-14 | ||
| K00962 (pnp, PNPT1) | maps-> | Purine metabolism | ||
| RNA degradation | ||||
| Pyrimidine metabolism | ||||
| InterPro domain | [2-48] IPR012162 | 3.8e-13 | Polyribonucleotide nucleotidyltransferase | |
| Orthology group | ||||
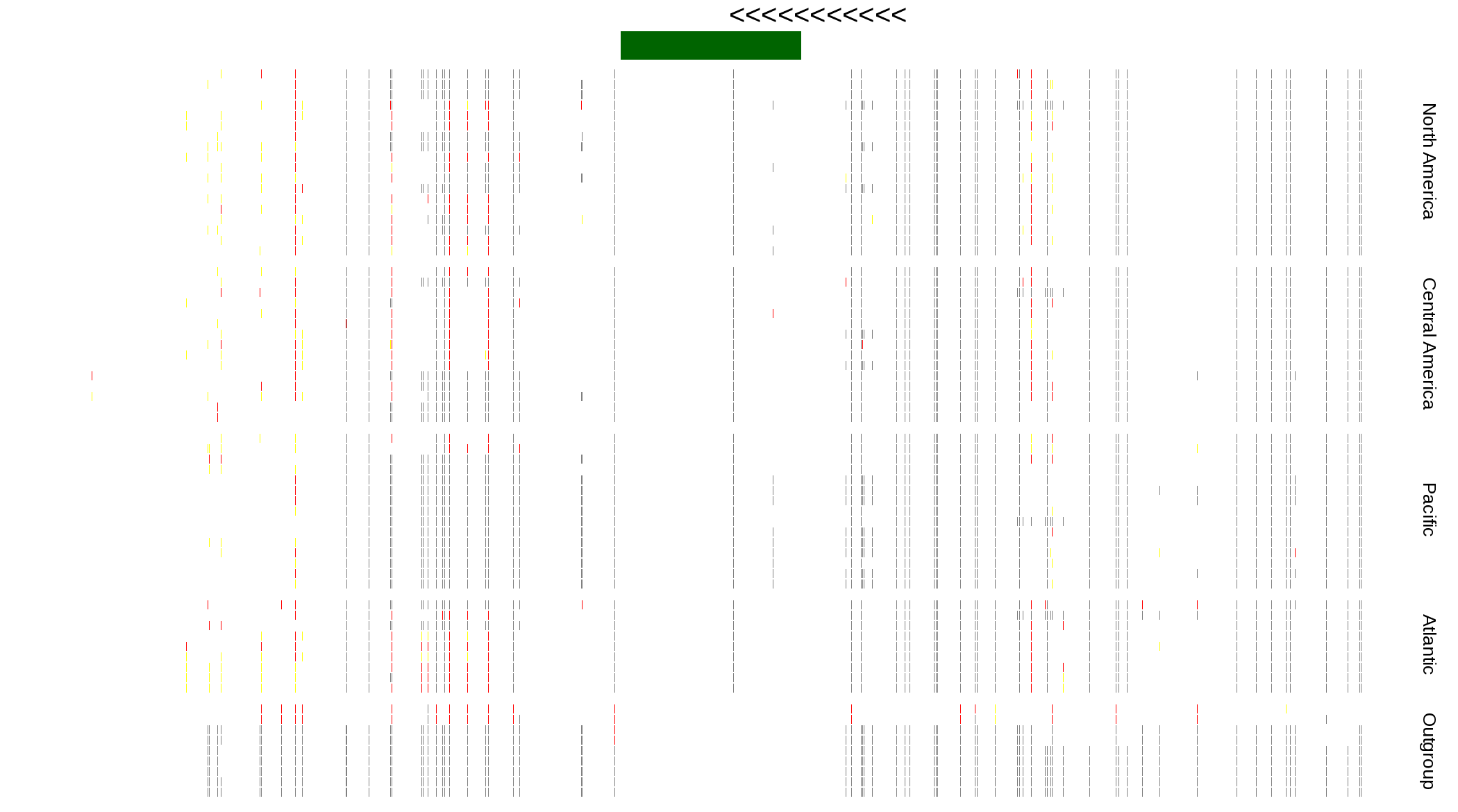
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204021-TA
ATGCACCCATCAGCGCTCGGCCTGACCGTGGGCTCGGAGATACAAGTCAAATACTTCGGTCGTGATCCCGTTTCCGGTCAAATGAGGCTCTCGAGGAAGGTTCTGACGTCACCTCCGCCGGGCATAGTCAGGAACCACGACAAGAGCTAG
>DPOGS204021-PA
MHPSALGLTVGSEIQVKYFGRDPVSGQMRLSRKVLTSPPPGIVRNHDKS-