| DPOGS204204 | ||
|---|---|---|
| Transcript | DPOGS204204-TA | 591 bp |
| Protein | DPOGS204204-PA | 196 aa |
| Genomic position | DPSCF300034 + 612072-613119 | |
| RNAseq coverage | 526x (Rank: top 24%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL004125 | 5e-101 | 87.24% | |
| Bombyx | BGIBMGA005162-TA | 5e-97 | 83.67% | |
| Drosophila | mRpL11-PA | 7e-75 | 64.29% | |
| EBI UniRef50 | UniRef50_Q9VFJ2 | 8e-73 | 64.29% | 39S ribosomal protein L11, mitochondrial n=18 Tax=Endopterygota RepID=RM11_DROME |
| NCBI RefSeq | XP_002070597.1 | 6e-76 | 65.31% | GK10947 [Drosophila willistoni] |
| NCBI nr blastp | % | |||
| NCBI nr blastx | % | |||
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005840 | 3.9e-61 | ribosome | |
| GO:0006412 | 3.9e-61 | translation | ||
| GO:0003735 | 3.9e-61 | structural constituent of ribosome | ||
| KEGG pathway | pmx:PERMA_1185 | 2e-26 | ||
| K02867 (RP-L11, rplK) | maps-> | Ribosome | ||
| InterPro domain | [24-158] IPR000911 | 3.9e-61 | Ribosomal protein L11 | |
| [88-157] IPR020783 | 1.2e-23 | Ribosomal protein L11, C-terminal domain | ||
| [26-82] IPR020784 | 4.6e-23 | Ribosomal protein L11, N-terminal | ||
| Orthology group | MCL13525 | Single-copy universal gene | ||
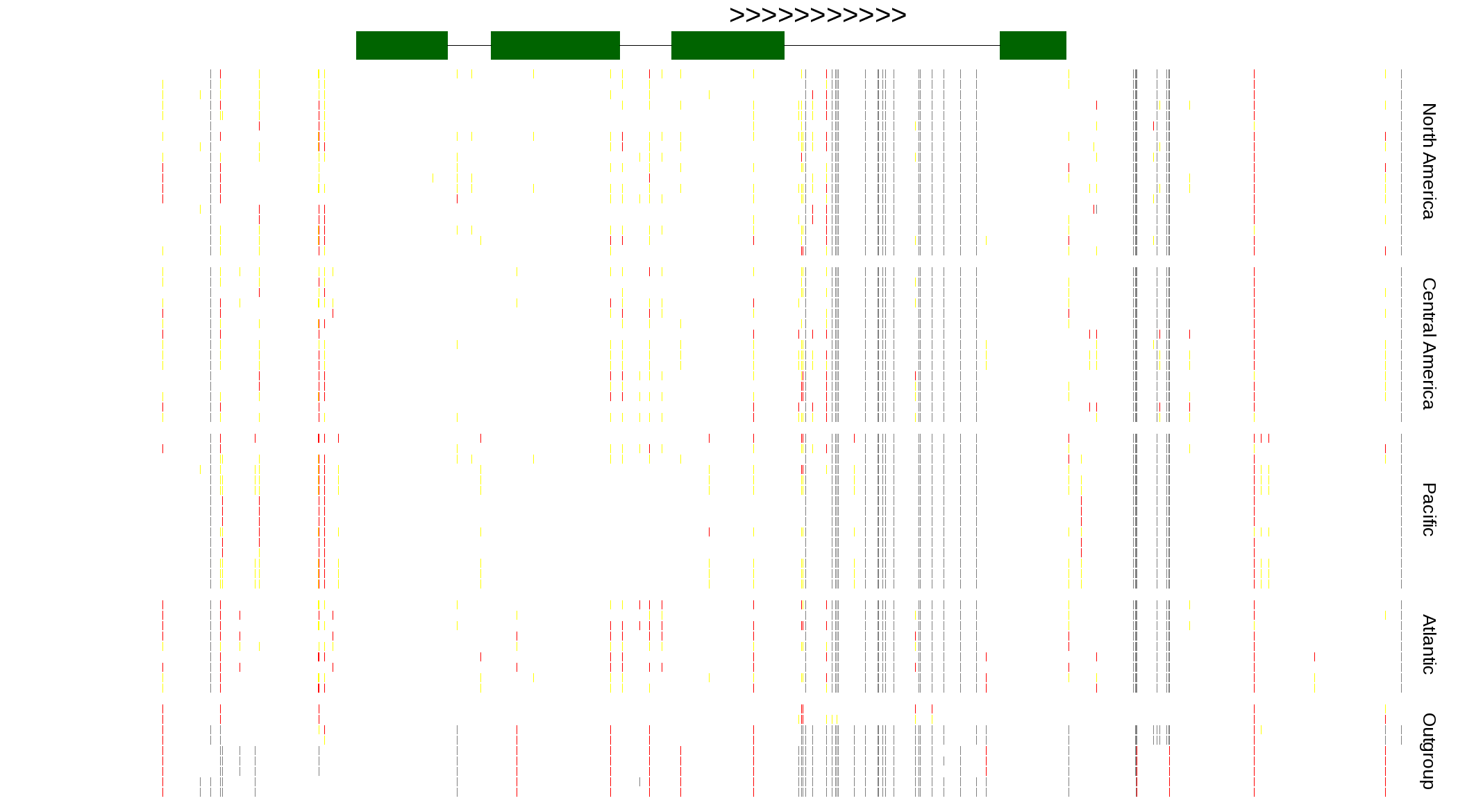
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204204-TA
ATGTCTAAGTCTGTGTCTAGATTGAAGTCAATGAAAAAAGTAGGTGACAAAATTATTCACTCCACTAAACTTGTAACCAATATTCCTGCGGGTATGGCTGCTGCAGGTCCTCCTTTAGGGCCTATGTTAGGACAGCGTAACATTAACATTGCAGCTTTTTGTAAAGACTTTAACGAGCGTACTGCTAGTATAAAACCTGGTATTCCATTACCCACTCGTGTGAAAGTGAATGCTGATAGATCATATCAATTAGTGATACATCAACCTCCCGCTAGTTATTTCCTGAAGCAAGCTGCTGGTGCTAGCCGAGGAGCTATGGAACCAGTCAAAGAGATATGCGGGAAAATAACACTTAAGCATATATATGAAATAGCAAAAATCAAGTCTCAAGATCCAACTCTCGAATGGAAACCATTGAAAGAAGTTTGCACCATGTTAATAGCTACGGCCAGAACATGTGGTATTGAAGTAGTCAAAGAATTAGATGCCAAGGAGTACGGTCAATTCCTTGAAGATAGGAAAATGATAATAGAGGAACAGAAGAAATTGTTACAAGAAAGAAGAGAAGCCAAAATGTTGAGGACTGCTTGA
>DPOGS204204-PA
MSKSVSRLKSMKKVGDKIIHSTKLVTNIPAGMAAAGPPLGPMLGQRNINIAAFCKDFNERTASIKPGIPLPTRVKVNADRSYQLVIHQPPASYFLKQAAGASRGAMEPVKEICGKITLKHIYEIAKIKSQDPTLEWKPLKEVCTMLIATARTCGIEVVKELDAKEYGQFLEDRKMIIEEQKKLLQERREAKMLRTA-