| DPOGS204652 | ||
|---|---|---|
| Transcript | DPOGS204652-TA | 492 bp |
| Protein | DPOGS204652-PA | 163 aa |
| Genomic position | DPSCF300170 - 605771-606922 | |
| RNAseq coverage | 0x (Rank: top 96%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL008363 | 5e-34 | 40.76% | |
| Bombyx | BGIBMGA007485-TA | 6e-42 | 45.86% | |
| Drosophila | beta4GalNAcTA-PA | 2e-42 | 45.81% | |
| EBI UniRef50 | UniRef50_E0VDW3 | 6e-42 | 47.74% | Putative uncharacterized protein n=1 Tax=Pediculus humanus corporis RepID=E0VDW3_PEDHC |
| NCBI RefSeq | XP_002424307.1 | 1e-42 | 47.74% | conserved hypothetical protein [Pediculus humanus corporis] |
| NCBI nr blastp | gi|242006954 | 2e-41 | 47.74% | conserved hypothetical protein [Pediculus humanus corporis] |
| NCBI nr blastx | gi|242006954 | 1e-41 | 47.74% | conserved hypothetical protein [Pediculus humanus corporis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016757 | 1.1e-66 | transferase activity, transferring glycosyl groups | |
| GO:0005975 | 1.1e-66 | carbohydrate metabolic process | ||
| KEGG pathway | cbr:CBG22165 | 4e-38 | ||
| K07968 (B4GALT3) | maps-> | Glycosphingolipid biosynthesis - lacto and neolacto series | ||
| Glycosaminoglycan biosynthesis - keratan sulfate | ||||
| N-Glycan biosynthesis | ||||
| InterPro domain | [1-158] IPR003859 | 1.1e-66 | Galactosyltransferase, metazoa | |
| Orthology group | ||||
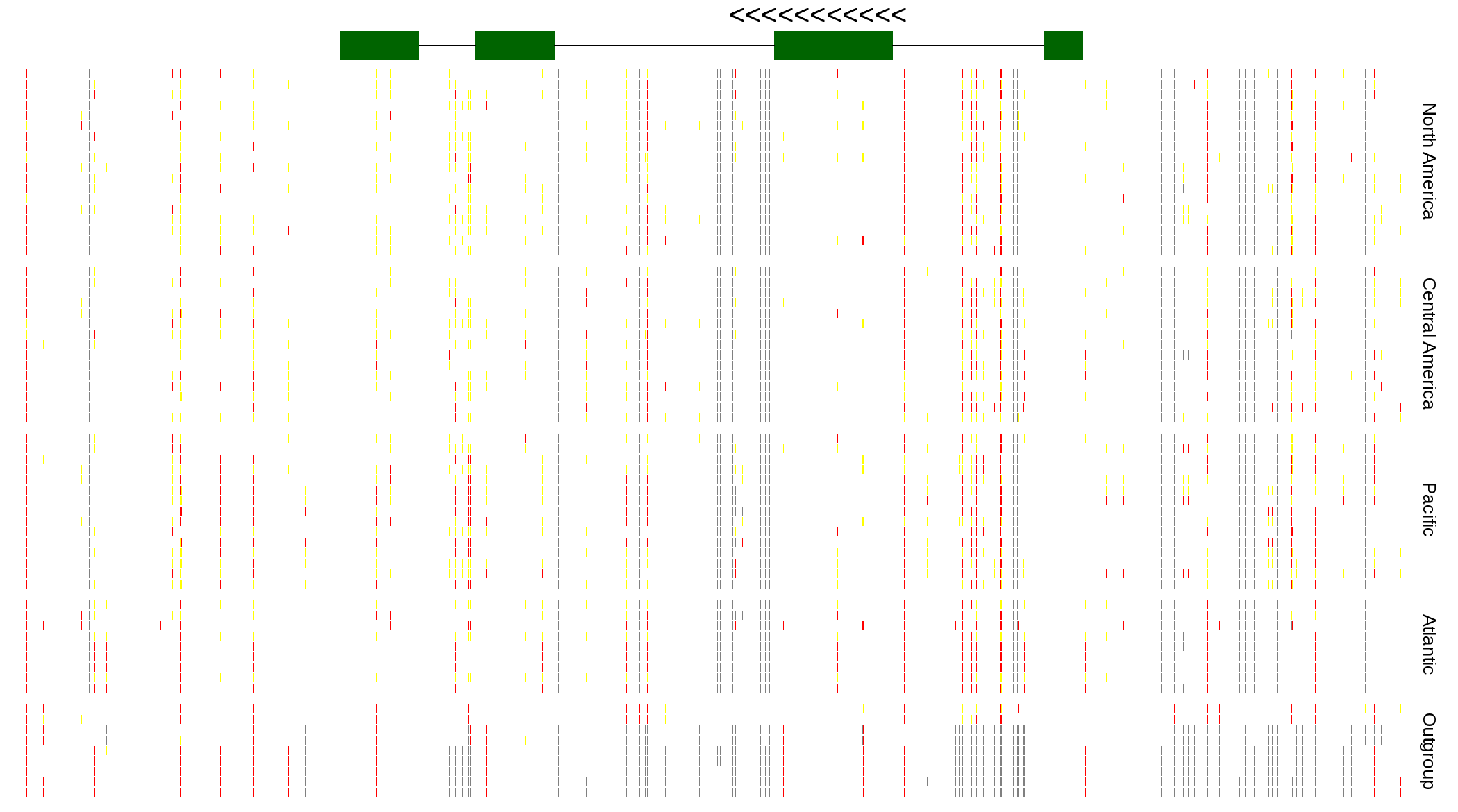
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204652-TA
ATGCATAAGTTCATGAAAAAGCAACTATTGGAATACCAAATTTTCCTGATAGAACAAATAGATAAGGACAGATTCAATAAAGGAGCATTATTCAACATAGGCTATACAGTATCACGAGTGTTTGGTAATTGGCATTGCTACGTCTTCCATGATGTAGATCTTCTCCCGATCGACGGCAGAGTATTGTACTCCTGTCCTCAACAACCCACTCACATGTCATCGGCGGTGGACTGTTATGATTTCAGACTGCCATATTATGGGTTATTTGGTGGTGTGACTATTCTAAAGCCTACTCATTACGAAGCTGTAAACGGTTATTCGAATTATTATTGGGATTGGGGGGCAGAAGACGACGATATGTACGCCAGATTGCAGGCACAAAAGTTCCCTGTGTTACATTATAGCTACTCATTAGGGAGATACGCAACTCTACCTCATGATCCAGCAAAAGAGAACTCCCAAAAGCAAGTATATCTGGTCATAAATACATAA
>DPOGS204652-PA
MHKFMKKQLLEYQIFLIEQIDKDRFNKGALFNIGYTVSRVFGNWHCYVFHDVDLLPIDGRVLYSCPQQPTHMSSAVDCYDFRLPYYGLFGGVTILKPTHYEAVNGYSNYYWDWGAEDDDMYARLQAQKFPVLHYSYSLGRYATLPHDPAKENSQKQVYLVINT-