| DPOGS204804 | ||
|---|---|---|
| Transcript | DPOGS204804-TA | 498 bp |
| Protein | DPOGS204804-PA | 165 aa |
| Genomic position | DPSCF300460 + 132697-133456 | |
| RNAseq coverage | 2x (Rank: top 91%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL013187 | 3e-77 | 88.02% | |
| Bombyx | BGIBMGA010833-TA | 3e-82 | 92.17% | |
| Drosophila | slou-PA | 2e-50 | 70.99% | |
| EBI UniRef50 | UniRef50_Q16J89 | 2e-52 | 66.27% | Nk homeobox protein n=2 Tax=Endopterygota RepID=Q16J89_AEDAE |
| NCBI RefSeq | XP_001663597.1 | 4e-53 | 66.27% | nk homeobox protein [Aedes aegypti] |
| NCBI nr blastp | gi|157135796 | 5e-52 | 66.27% | nk homeobox protein [Aedes aegypti] |
| NCBI nr blastx | gi|343168815 | 2e-56 | 69.82% | RT11964p1 [Drosophila melanogaster] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003677 | 4.6e-26 | DNA binding | |
| GO:0006355 | 4.6e-26 | regulation of transcription, DNA-dependent | ||
| GO:0043565 | 5e-24 | sequence-specific DNA binding | ||
| GO:0003700 | 5e-24 | sequence-specific DNA binding transcription factor activity | ||
| GO:0005515 | 2.3e-23 | protein binding | ||
| KEGG pathway | ||||
| InterPro domain | [18-99] IPR012287 | 4.6e-26 | Homeodomain-related | |
| [40-102] IPR001356 | 5e-24 | Homeobox | ||
| [39-113] IPR009057 | 2.3e-23 | Homeodomain-like | ||
| [62-73] IPR020479 | 4.8e-07 | Homeobox, eukaryotic | ||
| Orthology group | MCL16913 | Patchy | ||
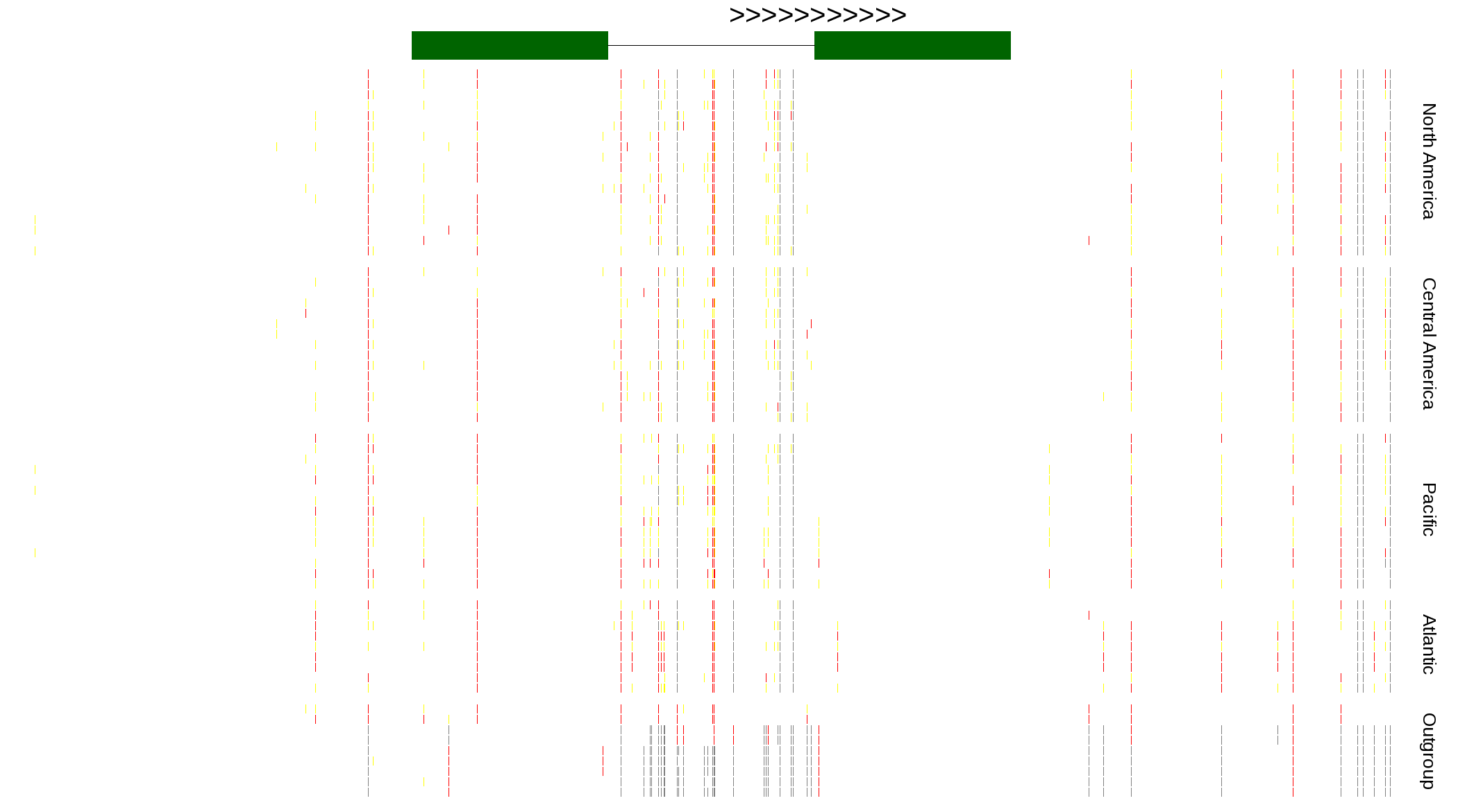
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204804-TA
ATGGATGATGAGATCTTAACACAATCAGATTTAGACGAAAATGGTGACCCAAAGAGCCCCACCAGCTCTAAGAAAGGGAAGAACGGTTCGTCCAGAGATCAGAAGGGAGGGACCAAGCCGAGACGAGCCAGAACAGCGTTCACGTACGAACAGCTGGTGTCTTTGGAGAATAAATTTAAGACTACTAGATATCTCTCCGTGTGCGAGCGACTTAATCTCGCTTTGAGCCTCAGTTTAACAGAAACCCAGGTCAAGATCTGGTTTCAAAATCGTCGTACGAAATGGAAAAAGCAGAACCCGGGTATGGACGTGAATAGTCCGACGGTGCCCCCTCCGCCGGCGGGGGCCTTTCCGTCGGGAGCTTATCCGGGGGGCCTTCTATACTCCCACGGGGTTCCATACCCCTTCACCGGGCCGGGGCCCTACGCCCCTTACTTCCACCATCTGGCTGCGAACCACCACCACAGCCACGGCTCCTTGGGACATTCCCATACGTGA
>DPOGS204804-PA
MDDEILTQSDLDENGDPKSPTSSKKGKNGSSRDQKGGTKPRRARTAFTYEQLVSLENKFKTTRYLSVCERLNLALSLSLTETQVKIWFQNRRTKWKKQNPGMDVNSPTVPPPPAGAFPSGAYPGGLLYSHGVPYPFTGPGPYAPYFHHLAANHHHSHGSLGHSHT-