| DPOGS204847 | ||
|---|---|---|
| Transcript | DPOGS204847-TA | 513 bp |
| Protein | DPOGS204847-PA | 170 aa |
| Genomic position | DPSCF300227 - 38601-39641 | |
| RNAseq coverage | 26x (Rank: top 77%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL012796 | 2e-09 | 35.45% | |
| Bombyx | BGIBMGA011740-TA | 1e-85 | 98.00% | |
| Drosophila | CG14950-PB | 2e-39 | 48.65% | |
| EBI UniRef50 | UniRef50_D6WGC8 | 1e-42 | 57.14% | Putative uncharacterized protein n=3 Tax=Tribolium castaneum RepID=D6WGC8_TRICA |
| NCBI RefSeq | XP_971304.2 | 2e-43 | 57.14% | PREDICTED: similar to CG16976 CG16976-PA [Tribolium castaneum] |
| NCBI nr blastp | gi|189235806 | 2e-42 | 57.14% | PREDICTED: similar to CG16976 CG16976-PA [Tribolium castaneum] |
| NCBI nr blastx | gi|270004703 | 9e-42 | 57.14% | hypothetical protein TcasGA2_TC010376 [Tribolium castaneum] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005515 | 1.6e-19 | protein binding | |
| KEGG pathway | dme:Dmel_CG11621 | 6e-11 | ||
| K00923 (E2.7.1.154, PIK3C2) | maps-> | Phosphatidylinositol signaling system | ||
| Inositol phosphate metabolism | ||||
| InterPro domain | [16-165] IPR008973 | 1.6e-19 | C2 calcium/lipid-binding domain, CaLB | |
| [53-126] IPR000008 | 1.8e-08 | C2 calcium-dependent membrane targeting | ||
| Orthology group | MCL19599 | Insect specific | ||
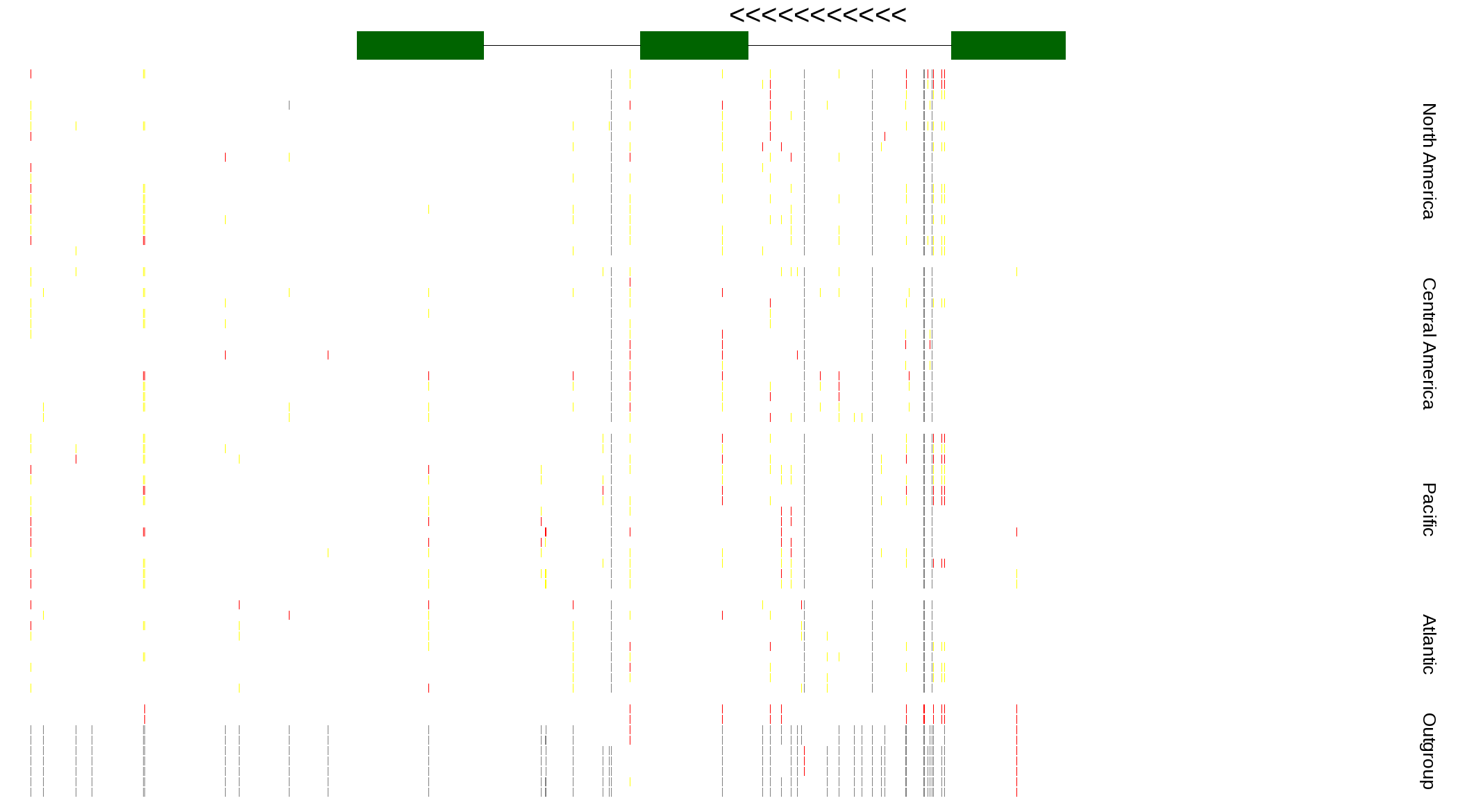
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS204847-TA
ATGTGGGCGGCGGTACCCGCAGCGGCGGCATGTGACGCGGTGGCAGACGACGATGACGACGTCCCTCTAGGTCCGGGACAACTGCCGCCGCGCAACGCTCATCTGCCGCCACTGCACGCTGAGATCAATATCAGCATCATCATGATCAAAGGACAACTTGAGCTAGAGGTATCTCATGCTCGTCGGGTTTATGGCGTTAATGGCGAGGTGCCTGACTCATACGTGAAATGCTACCTTCGCGATGGTGACAAGTGGTTGCACAAGCGTAAGACAAGAGTAATACGCCGCACCACTGAGCCACATTTCAAACAAACGCTTAAATATCAGGCAAGCGAGGCCTTAGGTCGTACGCTGGTAGTGATGCTGTGGCAACGATGCGGGGGATTTGAACATAACCTTGCCCTGGGAGGTGCGGAAATATGTCTCGACAAACTGTCACTGCCGCAGCGCACGTATGGCTGGTACCCTCTGTTCCCAGCCACCTCGCTCGCCGCTGACGAGTCGCCCGATTGA
>DPOGS204847-PA
MWAAVPAAAACDAVADDDDDVPLGPGQLPPRNAHLPPLHAEINISIIMIKGQLELEVSHARRVYGVNGEVPDSYVKCYLRDGDKWLHKRKTRVIRRTTEPHFKQTLKYQASEALGRTLVVMLWQRCGGFEHNLALGGAEICLDKLSLPQRTYGWYPLFPATSLAADESPD-