| DPOGS205467 | ||
|---|---|---|
| Transcript | DPOGS205467-TA | 675 bp |
| Protein | DPOGS205467-PA | 224 aa |
| Genomic position | DPSCF300166 - 70566-72786 | |
| RNAseq coverage | 0x (Rank: top 97%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL013003 | 3e-37 | 40.08% | |
| Bombyx | BGIBMGA008278-TA | 4e-34 | 39.19% | |
| Drosophila | CG13430-PB | 2e-17 | 27.16% | |
| EBI UniRef50 | UniRef50_A5CG73 | 1e-31 | 36.73% | Chymotrypsinogen-like protein 3 n=9 Tax=Obtectomera RepID=A5CG73_MANSE |
| NCBI RefSeq | NP_001103386.1 | 8e-19 | 27.54% | cocoonase [Bombyx mori] |
| NCBI nr blastp | gi|151199948 | 4e-37 | 37.55% | chymotrypsin-like protease C1 [Heliothis virescens] |
| NCBI nr blastx | gi|171740887 | 6e-33 | 45.30% | trypsin [Helicoverpa armigera] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003824 | 4.6e-45 | catalytic activity | |
| GO:0004252 | 1.6e-25 | serine-type endopeptidase activity | ||
| GO:0006508 | 1.6e-25 | proteolysis | ||
| KEGG pathway | ||||
| InterPro domain | [25-218] IPR009003 | 4.6e-45 | Peptidase cysteine/serine, trypsin-like | |
| [35-213] IPR001254 | 1.6e-25 | Peptidase S1/S6, chymotrypsin/Hap | ||
| Orthology group | MCL23321 | Specific divergent | ||
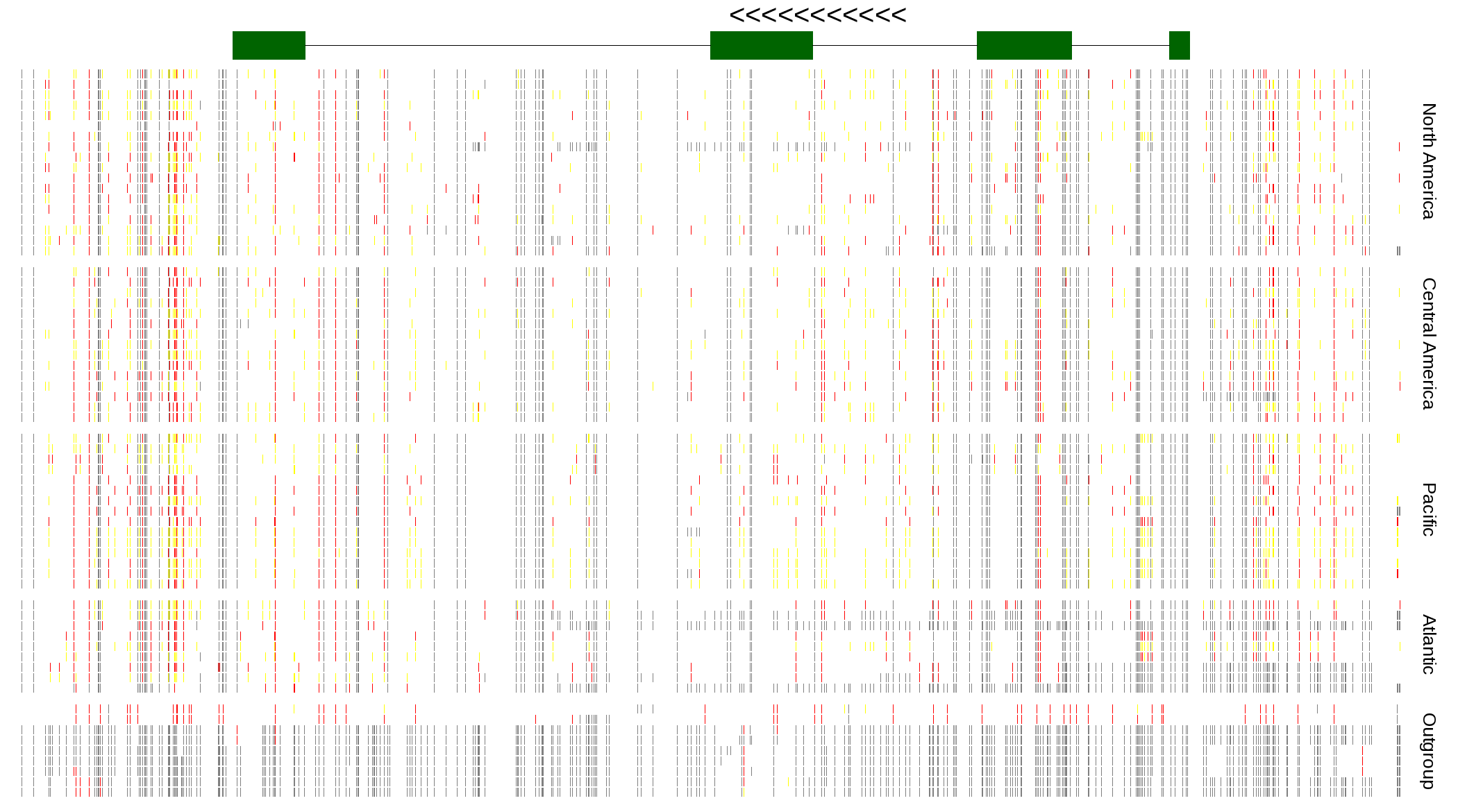
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS205467-TA
ATGTTTCAGGCTGGGTGTTGGTTTATATCGCTCTTAGTGGGAAGCATAGGTTTGCCCTTAAGTGGTGGATACACATCCGGTTACTTCGGCGATATAAAAGCTCGGGTCATCAATGGTTACGAAGCAAATCTCGTCCCTCACATGGTAGCTCTGACTGTCGGTGAAGATAGTAAAATCCTCGTATGTGGTGCCTCCCTTATTACTAAACGCCATGTACTCACAGCCGCTCACTGCATTGACGCTTTTATAGATAAAGGAAAATTGAGCAGGTCTCTTAAAGGAATTGTTGGGACAAATGATTGGAACAAGGGCGGAACTCATTACACCTTTTCTGGCAACATTACTCATCCTGAATGGGATGCAAAAAATGTCAATAATGATATTGGTATACTTATCACGTCACAACCGGTGACACTTAATAAATATGTACAGATCATTACCCTTAATTTTCAATTTATTGGTGGTAACGTCGCTGCTATTATCAACGGTTGGGGACAATTTGAAAAGGGTGATTCGGGTAGCCCGTTAATCCTACGAGATAATAGACAGCAAATTGGCGTCGTGTCCTGGGGTGTCAACCCCAGTGCATCCGGATATCCTGATGTGTACGGTAGAATCAGTGCATACAAATCTTGGATTAAACAGAGCGTACGATTAAGAATACGAGTAAATTAA
>DPOGS205467-PA
MFQAGCWFISLLVGSIGLPLSGGYTSGYFGDIKARVINGYEANLVPHMVALTVGEDSKILVCGASLITKRHVLTAAHCIDAFIDKGKLSRSLKGIVGTNDWNKGGTHYTFSGNITHPEWDAKNVNNDIGILITSQPVTLNKYVQIITLNFQFIGGNVAAIINGWGQFEKGDSGSPLILRDNRQQIGVVSWGVNPSASGYPDVYGRISAYKSWIKQSVRLRIRVN-