| DPOGS205623 | ||
|---|---|---|
| Transcript | DPOGS205623-TA | 540 bp |
| Protein | DPOGS205623-PA | 179 aa |
| Genomic position | DPSCF300023 - 530816-531967 | |
| RNAseq coverage | 2117x (Rank: top 6%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006212 | 3e-92 | 96.09% | |
| Bombyx | BGIBMGA001136-TA | 4e-55 | 94.29% | |
| Drosophila | Spase22-23-PA | 2e-77 | 73.74% | |
| EBI UniRef50 | UniRef50_G6CUW8 | 5e-100 | 100.00% | Signal peptidase complex subunit 3 n=2 Tax=Coelomata RepID=G6CUW8_DANPL |
| NCBI RefSeq | NP_001091763.1 | 4e-89 | 94.41% | signal peptidase complex subunit 3 [Bombyx mori] |
| NCBI nr blastp | gi|148298671 | 6e-88 | 94.41% | signal peptidase complex subunit 3 [Bombyx mori] |
| NCBI nr blastx | gi|148298671 | 1e-94 | 94.41% | signal peptidase complex subunit 3 [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0008233 | 8e-89 | peptidase activity | |
| GO:0016021 | 8e-89 | integral to membrane | ||
| GO:0006465 | 8e-89 | signal peptide processing | ||
| GO:0005787 | 8e-89 | signal peptidase complex | ||
| KEGG pathway | aag:AaeL_AAEL000947 | 5e-77 | ||
| K01423 (E3.4.-.-) | maps-> | Biotin metabolism | ||
| Lysine degradation | ||||
| InterPro domain | [2-180] IPR007653 | 8e-89 | Signal peptidase 22kDa subunit | |
| Orthology group | MCL14670 | Multiple-copy universal gene | ||
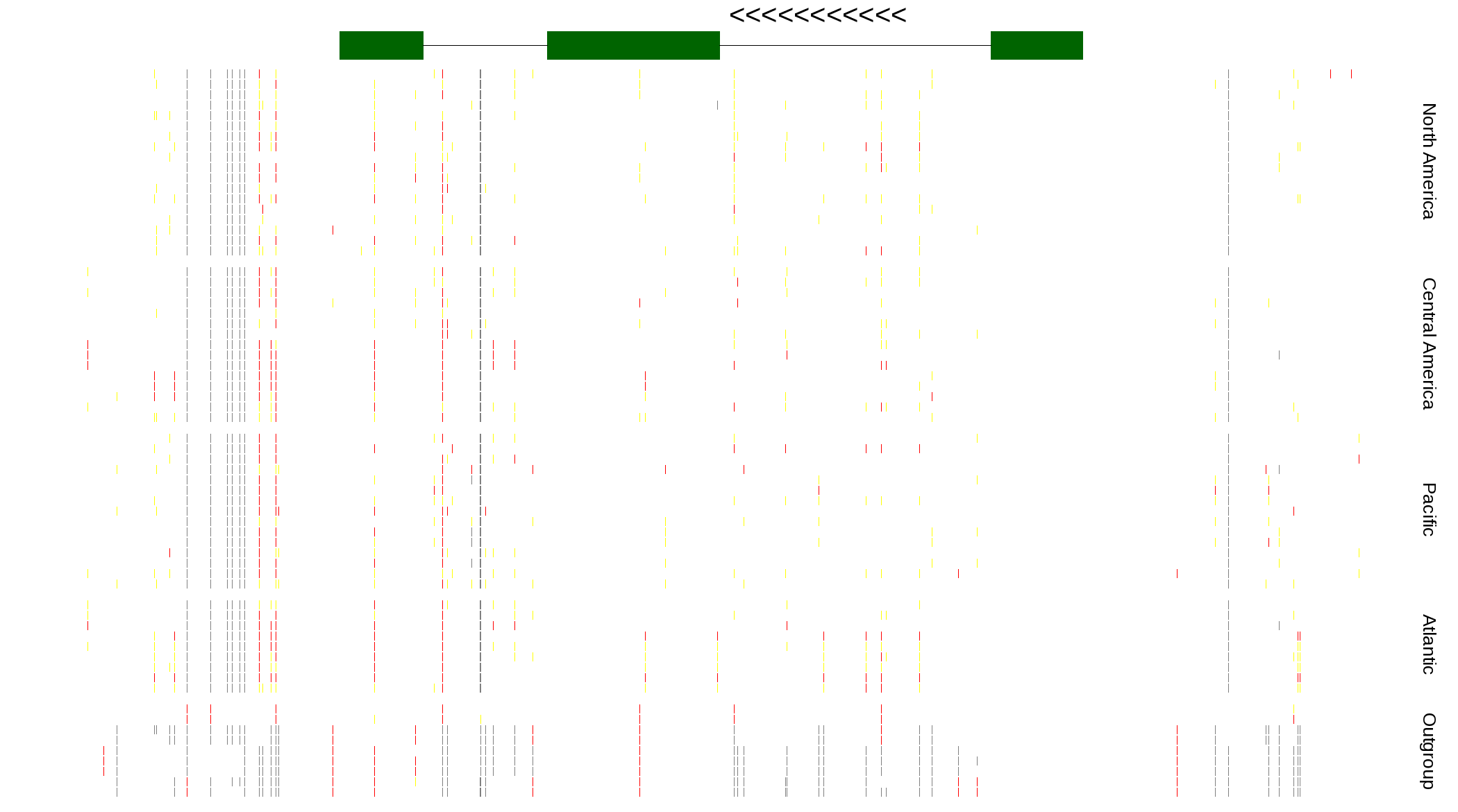
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS205623-TA
ATGTATTCCGTTATAACCAGAGTCAACGCGATATTAACGTACACACTAAGTGTTCTAGCATGTCTTACATTTTTGTGTTTTCTTTCCACTCTAACAGTGGATTATAGAACTACTGCTCAGATGAATACCGTAAAAGTTGTGGTGAAGAATGTTCCAGACTATGGTGCTTCAAGGGAAAGAAATGATCTTGGTTACTTAACATTTGATCTCAAAACAGATTTATCCAACCTATTCAACTGGAATGTTAAACAGTTGTTTTTGTACCTAACAGCCGAGTATATCACACCAAACAATGAACTAAACCAAGTTGTACTATGGGACAAGATTATACTCAGAGGAGAGAATGCATTGTTAGATTTTAAAAACATGAACACTAAATACTATTTCTGGGATGACGGGAATGGTTTGAAAGGTCACAACAACGTCACATTAACCCTCTCCTGGAATATTATACCTAATGCCGGCCTGTTGCCAAACATCCAGGCCGTTGGTTTGCATTCATTCAAGTTTCCTACAGAATACACTCAAACCAGAGTATGA
>DPOGS205623-PA
MYSVITRVNAILTYTLSVLACLTFLCFLSTLTVDYRTTAQMNTVKVVVKNVPDYGASRERNDLGYLTFDLKTDLSNLFNWNVKQLFLYLTAEYITPNNELNQVVLWDKIILRGENALLDFKNMNTKYYFWDDGNGLKGHNNVTLTLSWNIIPNAGLLPNIQAVGLHSFKFPTEYTQTRV-