| DPOGS205650 | ||
|---|---|---|
| Transcript | DPOGS205650-TA | 597 bp |
| Protein | DPOGS205650-PA | 198 aa |
| Genomic position | DPSCF300023 + 164799-165395 | |
| RNAseq coverage | 9x (Rank: top 85%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL004170 | 5e-61 | 69.42% | |
| Bombyx | BGIBMGA001000-TA | 9e-47 | 64.40% | |
| Drosophila | sc-PA | 1e-19 | 54.67% | |
| EBI UniRef50 | UniRef50_A6N868 | 1e-47 | 63.11% | Achaete-scute-like protein ASH3 n=4 Tax=Obtectomera RepID=A6N868_BOMMO |
| NCBI RefSeq | NP_001098694.1 | 3e-48 | 63.11% | achaete-scute-like protein ASH3 [Bombyx mori] |
| NCBI nr blastp | gi|157412306 | 5e-47 | 63.11% | achaete-scute-like protein ASH3 [Bombyx mori] |
| NCBI nr blastx | gi|157412306 | 2e-58 | 64.04% | achaete-scute-like protein ASH3 [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003677 | 4.1e-30 | DNA binding | |
| GO:0005634 | 2.6e-20 | nucleus | ||
| GO:0006355 | 2.6e-20 | regulation of transcription, DNA-dependent | ||
| KEGG pathway | ||||
| InterPro domain | [36-118] IPR015660 | 4.1e-30 | Achaete-scute transcription factor-related | |
| [39-114] IPR011598 | 2.6e-20 | Helix-loop-helix DNA-binding | ||
| [52-115] IPR001092 | 1.6e-18 | Helix-loop-helix DNA-binding domain | ||
| Orthology group | MCL26616 | Lepidoptera specific | ||
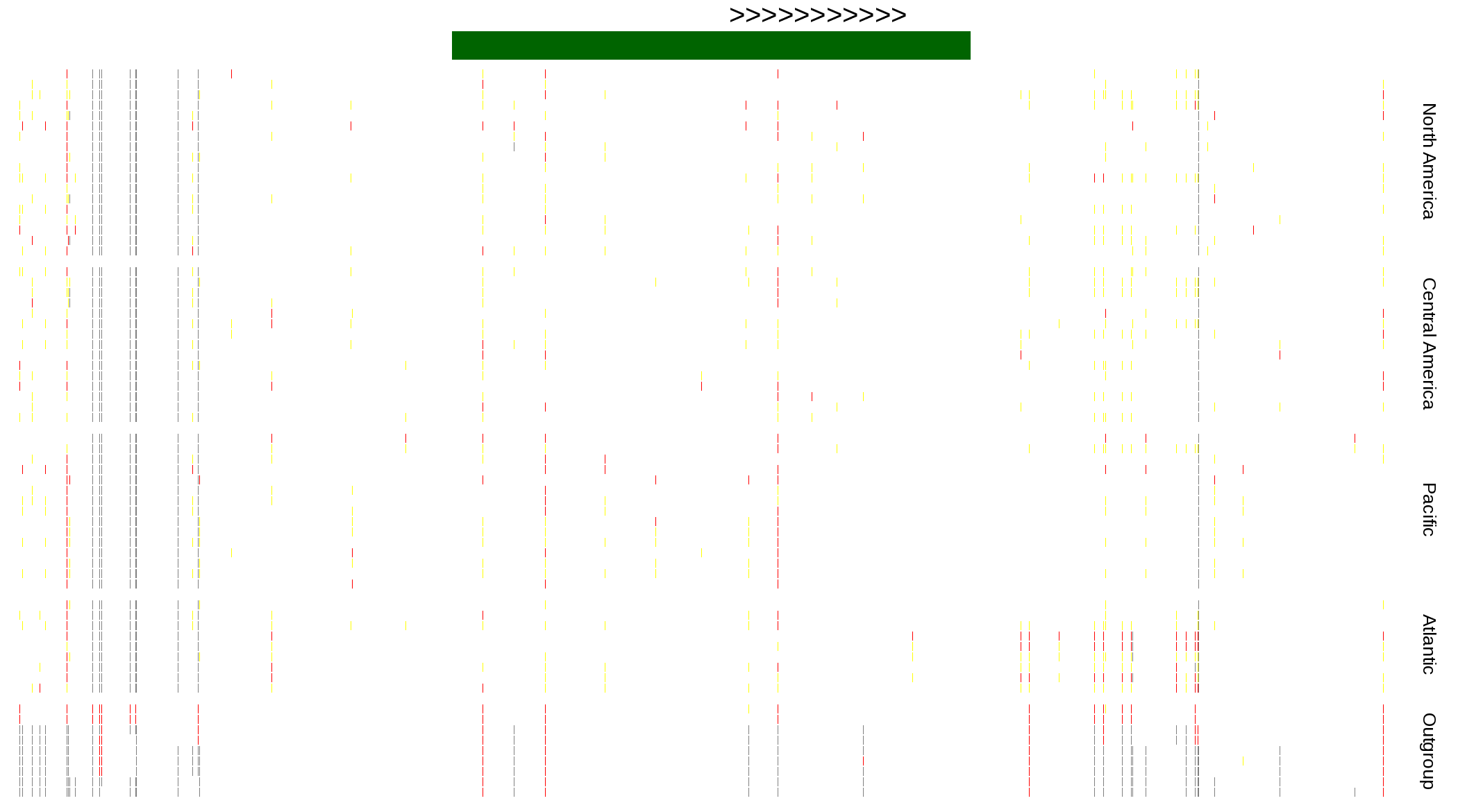
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS205650-TA
ATGATTCAAACCAAGAACTATACTGGCTTTTACAATAACGAGATCCCAAAATTCGTTCCAATCGCTCCCATGGTGGATCAAGAAATGGATTGCACAGTGCGAAAATACAATTACAAAAATAATTCCGGACAAGCAGCATCTATTGCGAGGCGCAATGCAAGAGAGAGGAATCGCGTTAAACAAGTTAATGATGGTTTTAACGCTTTAAGAAAAAGATTACCCGCTGCCGTTGTGAATGCATTGTCTGGAGGCGCTCGTCGGGGATCCGGCAAGAAATTGAGCAAAGTCGATACATTACGAATGGTGGTTGAGTACATTAAATATTTGGAGAATTTAATCGACGAAAGTGATGCTTCATTGGGTGTTACCAAAGAACCTATGGATACATCTGGAATTGATGAGGGAATTTTTGGAAGAACATCACCGTACTCTGACTCTGTACCATCGCCTGCTAACTCTGAATGTTCATCGGGGGTGTCTTCAAGCTATTCAAATGATCATTACCAAATTAACACTCACAACAATTACTATAGTGAAGTAAACGCAGTAAATGATAGTGAATTATTAGATGCGATCACTTGGTGGCAGCAAAAGTAA
>DPOGS205650-PA
MIQTKNYTGFYNNEIPKFVPIAPMVDQEMDCTVRKYNYKNNSGQAASIARRNARERNRVKQVNDGFNALRKRLPAAVVNALSGGARRGSGKKLSKVDTLRMVVEYIKYLENLIDESDASLGVTKEPMDTSGIDEGIFGRTSPYSDSVPSPANSECSSGVSSSYSNDHYQINTHNNYYSEVNAVNDSELLDAITWWQQK-