| DPOGS205699 | ||
|---|---|---|
| Transcript | DPOGS205699-TA | 381 bp |
| Protein | DPOGS205699-PA | 126 aa |
| Genomic position | DPSCF300250 - 19045-19425 | |
| RNAseq coverage | 335x (Rank: top 34%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL014801 | 2e-42 | 66.41% | |
| Bombyx | BGIBMGA009834-TA | 2e-41 | 65.62% | |
| Drosophila | CG15891-PA | 2e-06 | 31.08% | |
| EBI UniRef50 | UniRef50_UPI00015B4E7F | 4e-10 | 32.59% | UPI00015B4E7F related cluster n=1 Tax=unknown RepID=UPI00015B4E7F |
| NCBI RefSeq | XP_001607781.1 | 6e-11 | 32.59% | PREDICTED: similar to CG15891-PA [Nasonia vitripennis] |
| NCBI nr blastp | gi|156544968 | 2e-09 | 32.59% | PREDICTED: hypothetical protein LOC100114386 [Nasonia vitripennis] |
| NCBI nr blastx | gi|156544968 | 3e-11 | 32.84% | PREDICTED: hypothetical protein LOC100114386 [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ||||
| Orthology group | MCL16352 | Insect specific | ||
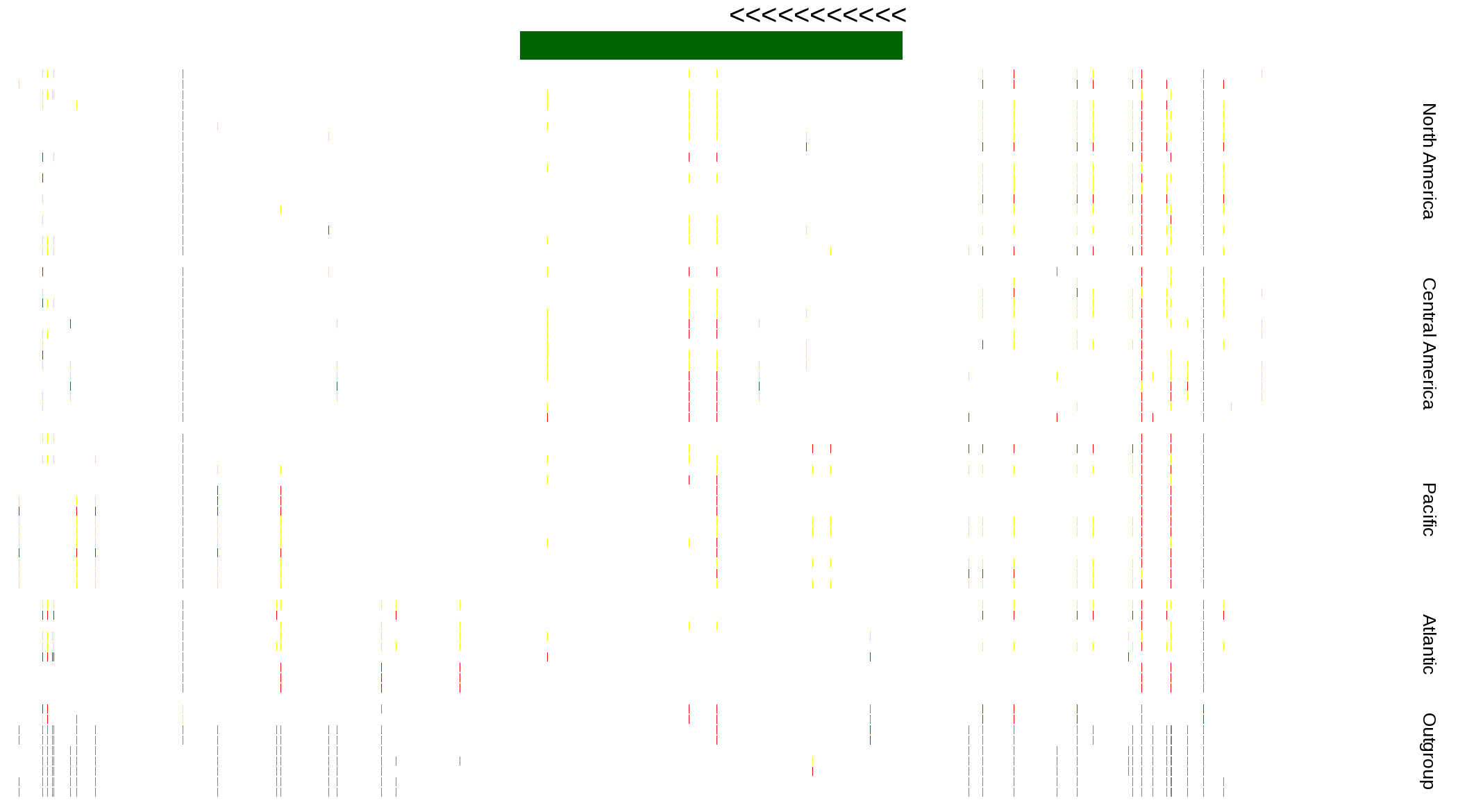
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS205699-TA
ATGTTTTCGTCATTTTTTGGAAAACGACGGTCATCACCCGTGGAAGATGAAACTCCACAAATTCCTGGACCGAAGTCAGATGACGGTTTCGTTATTGTAGATCAGACGTCTCCAAGAGGTAGCTTATATCCAAGTATCAGTGGGTCTGTGTATCCACAGCGACCGGCTCCACCGCCTCCAGTGGTCCCAAAACCGGAACAATGCTTCCATTACCTACAGGGAGTCCCATTTACACTGTCCCGTGAGTTACGGATGACAACAAATAAGGATGCAATAGCTACTGAAATCGGAGATATTCTAGCATTCCTCTCCAACAAGTTAAATGTGAACGACTATAACTATGATTTTTCTGTAGAAAAAAGTGTTTTGAAAGAATATTAA
>DPOGS205699-PA
MFSSFFGKRRSSPVEDETPQIPGPKSDDGFVIVDQTSPRGSLYPSISGSVYPQRPAPPPPVVPKPEQCFHYLQGVPFTLSRELRMTTNKDAIATEIGDILAFLSNKLNVNDYNYDFSVEKSVLKEY-