| DPOGS206233 | ||
|---|---|---|
| Transcript | DPOGS206233-TA | 291 bp |
| Protein | DPOGS206233-PA | 96 aa |
| Genomic position | DPSCF300334 + 109601-110830 | |
| RNAseq coverage | 0x (Rank: top 97%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL011234 | 3e-29 | 71.43% | |
| Bombyx | BGIBMGA009744-TA | 4e-33 | 77.78% | |
| Drosophila | CG10585-PA | 1e-11 | 59.57% | |
| EBI UniRef50 | UniRef50_UPI0000E49D87 | 6e-10 | 68.29% | UPI0000E49D87 related cluster n=2 Tax=unknown RepID=UPI0000E49D87 |
| NCBI RefSeq | XP_001190717.1 | 9e-11 | 68.29% | PREDICTED: similar to Prenyl (decaprenyl) diphosphate synthase, subunit 2 [Strongylocentrotus purpuratus] |
| NCBI nr blastp | gi|47220118 | 2e-10 | 63.64% | unnamed protein product [Tetraodon nigroviridis] |
| NCBI nr blastx | gi|348515925 | 5e-10 | 68.29% | PREDICTED: decaprenyl-diphosphate synthase subunit 2-like [Oreochromis niloticus] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | dre:436624 | 4e-11 | ||
| K12505 (PDSS2) | maps-> | Terpenoid backbone biosynthesis | ||
| InterPro domain | [30-89] IPR017446 | 4.3e-19 | Polyprenyl synthetase-related | |
| Orthology group | ||||
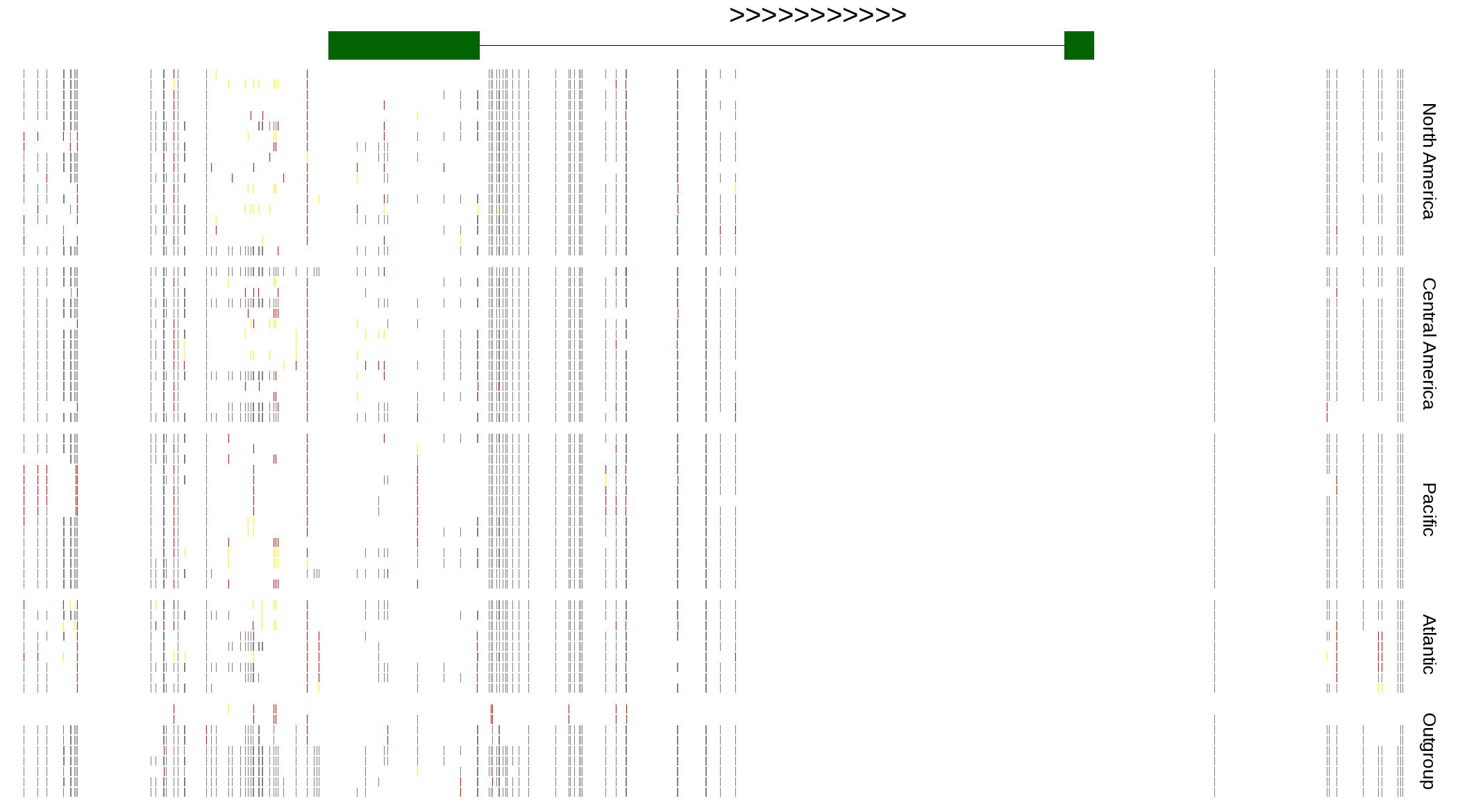
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206233-TA
ATGGCTCTACTGACTCGTGTGTTGAGGCGAGCGACGCCTTACCTCCACCACACCAGGCCGGAGTCCACAGTGGCCAGTTTCACCAAGGATGAGATCCTGATACTGCTGAGACCACCTTTCACGAACTGGAGCAACATCATCCGTGAGGCGGAGAAAGTTGTCGGCTATCCCACTTCCTTCATCAACCTGAGATGTCTCCTCAGCGACGAGTTCTCCAACCTGGCTCTGTACTTAAGGAAGCTGGATTCTAGTATCGTCCTCACAGTAAACTTGCAAACTGTCGGTTCTTAG
>DPOGS206233-PA
MALLTRVLRRATPYLHHTRPESTVASFTKDEILILLRPPFTNWSNIIREAEKVVGYPTSFINLRCLLSDEFSNLALYLRKLDSSIVLTVNLQTVGS-