| DPOGS206592 | ||
|---|---|---|
| Transcript | DPOGS206592-TA | 249 bp |
| Protein | DPOGS206592-PA | 82 aa |
| Genomic position | DPSCF300048 - 1729531-1731980 | |
| RNAseq coverage | 0x (Rank: top 97%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | % | |||
| Bombyx | BGIBMGA008324-TA | 3e-33 | 75.32% | |
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_B0X935 | 5e-07 | 35.62% | Hormone-sensitive lipase n=2 Tax=Culicinae RepID=B0X935_CULQU |
| NCBI RefSeq | XP_001866157.1 | 8e-08 | 35.62% | hormone-sensitive lipase [Culex quinquefasciatus] |
| NCBI nr blastp | gi|170061257 | 2e-06 | 35.62% | hormone-sensitive lipase [Culex quinquefasciatus] |
| NCBI nr blastx | gi|170061257 | 3e-06 | 35.62% | hormone-sensitive lipase [Culex quinquefasciatus] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016042 | 1.2e-08 | lipid catabolic process | |
| GO:0008203 | 1.2e-08 | cholesterol metabolic process | ||
| GO:0016298 | 1.2e-08 | lipase activity | ||
| KEGG pathway | ||||
| InterPro domain | [26-76] IPR010468 | 1.2e-08 | Hormone-sensitive lipase, N-terminal | |
| Orthology group | ||||
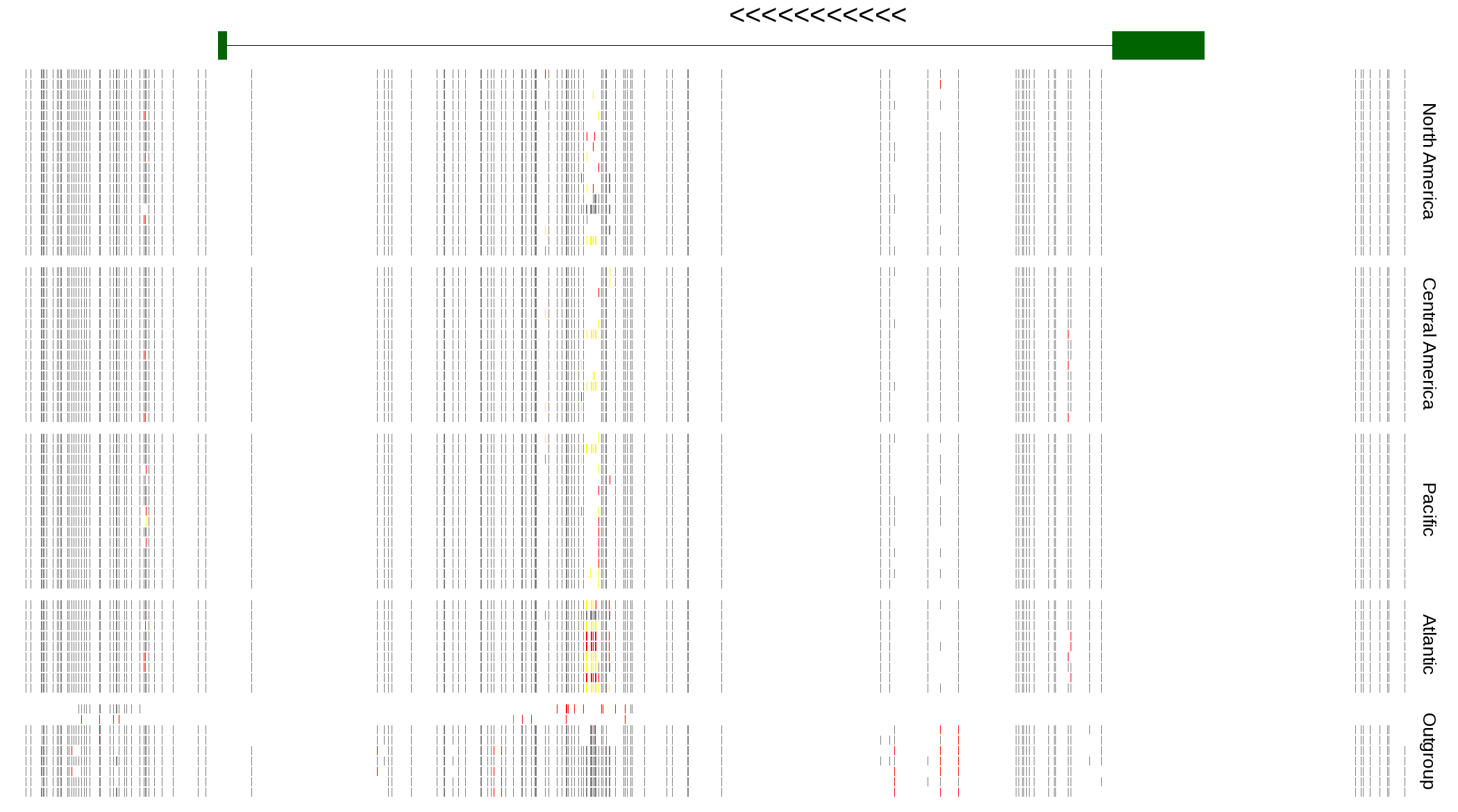
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206592-TA
ATGGAATTCGCGGAGCCCCCTGCGAAGGCATCATTATGCTGTGAGGATACTCCCGAAGGTTCTCCGCCGACGTATGCAATGTATGAGGCTTTAAAGGAATCGTGTCAGAATAATGCTAGTTATTTTCAACCTGATGACAGTGAGAATGGACAGCGATTGTACCAGGGCTTCATGACACTGATAGATCATATAGATACAGTTTGGCCTCTCGTGGATCACGTTCGAAAGGCCCTAGAAGTACTTCCATAG
>DPOGS206592-PA
MEFAEPPAKASLCCEDTPEGSPPTYAMYEALKESCQNNASYFQPDDSENGQRLYQGFMTLIDHIDTVWPLVDHVRKALEVLP-