| DPOGS206605 | ||
|---|---|---|
| Transcript | DPOGS206605-TA | 633 bp |
| Protein | DPOGS206605-PA | 210 aa |
| Genomic position | DPSCF300048 - 1273313-1274180 | |
| RNAseq coverage | 224x (Rank: top 44%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL008839 | 7e-96 | 79.26% | |
| Bombyx | % | |||
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_E9GF16 | 6e-35 | 40.29% | Putative uncharacterized protein n=1 Tax=Daphnia pulex RepID=E9GF16_DAPPU |
| NCBI RefSeq | XP_001605788.1 | 1e-43 | 44.50% | PREDICTED: similar to motile sperm domain containing 1 [Nasonia vitripennis] |
| NCBI nr blastp | gi|340713182 | 2e-44 | 43.46% | PREDICTED: motile sperm domain-containing protein 1-like isoform 1 [Bombus terrestris] |
| NCBI nr blastx | gi|380015472 | 2e-43 | 43.54% | PREDICTED: motile sperm domain-containing protein 1-like [Apis florea] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005198 | 8.2e-13 | structural molecule activity | |
| KEGG pathway | ||||
| InterPro domain | [2-140] IPR008962 | 3.6e-17 | PapD-like | |
| [17-89] IPR000535 | 8.2e-13 | Major sperm protein | ||
| Orthology group | MCL17981 | Patchy | ||
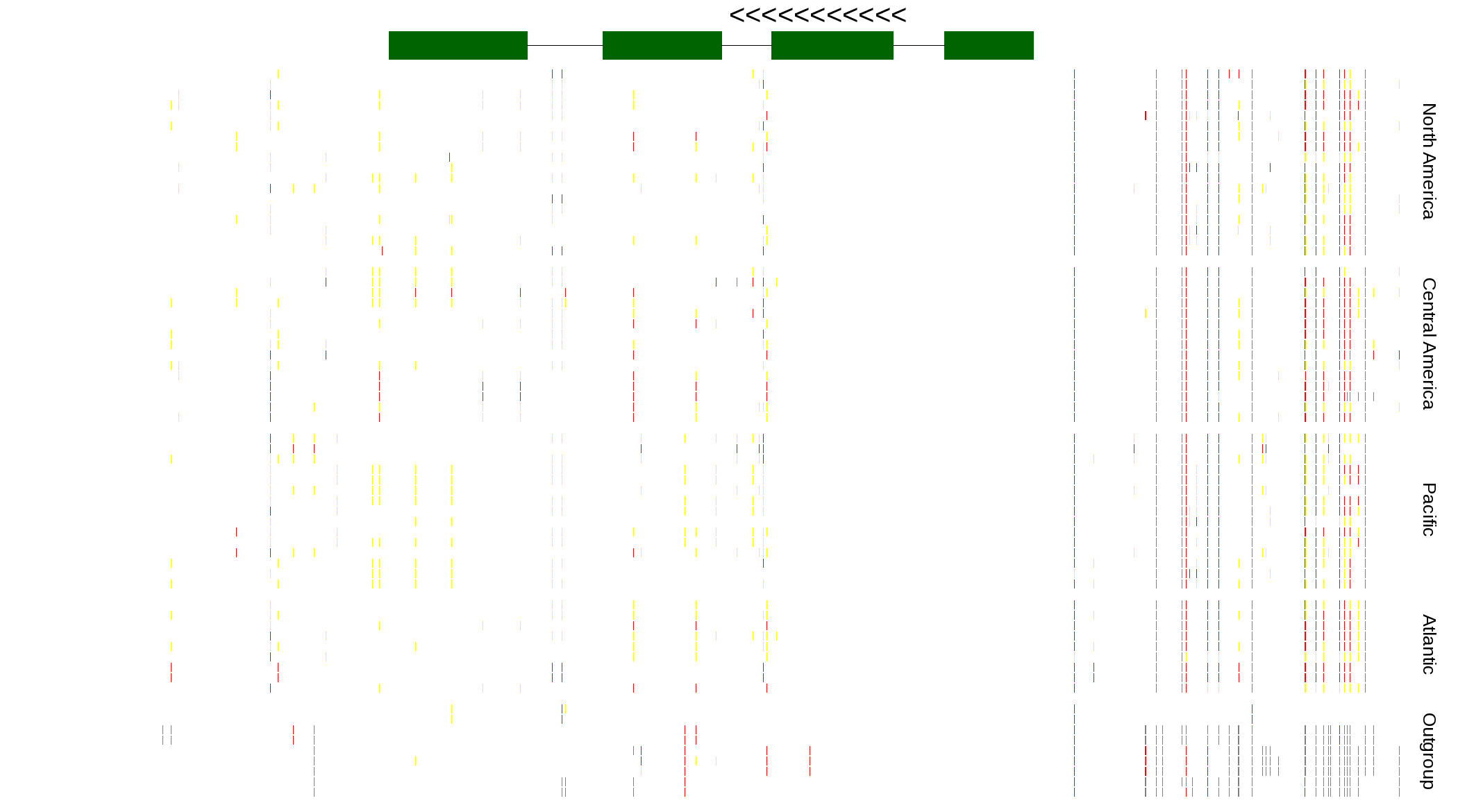
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206605-TA
ATGAAGAATTTCCCCGTGTTTGTCTTTCCAGTTTCCTTGGAGTTCTATTTAAACGCGAGACATACCCATAAACAGTTGTTAACTGTGTATAATCCATACGAGTTTTCTGTTAATTTCAAAGTATTATGTACATCTCCAAACAAATTCACTGTAATAGACCCAGAAGGAGTTATAGCTCCTCAAAGTTGTATCGATATCGTAGTTAGATATTCGCAGCCGTCTATCTCGCATTGCAATTCCGCTGAAAAGTTTCGAATAACAATGTATGATCGAAATACCCAACAGGCATTGGGTAAACGGGATATTCCAACCAAACTCATCGAAGGTGAACCATGCAACATCAATCAAGAAAGTTTGGGAGATAATTTTCATCCACTTACTACAAAACCAGCACCAATGAGAACGGAGGAACATTCTCGAGTCATATGTAATCATTCACACCACAGAGACCAACCCATCAATGTTGTGGCGGTGGCAGTGAGTATATGTTGTATAGCAGCCCTACTTTTGCCAACACAACCAGAACAGGTGCTGAAATCACAGTGGCCGGAGTGGTTGCATATCGGGTCTAATCTTAAGTTAGTATTTTCATTTGTCCTAGGACTTGTTAGCATGCTAGTATTAAGACCTTGA
>DPOGS206605-PA
MKNFPVFVFPVSLEFYLNARHTHKQLLTVYNPYEFSVNFKVLCTSPNKFTVIDPEGVIAPQSCIDIVVRYSQPSISHCNSAEKFRITMYDRNTQQALGKRDIPTKLIEGEPCNINQESLGDNFHPLTTKPAPMRTEEHSRVICNHSHHRDQPINVVAVAVSICCIAALLLPTQPEQVLKSQWPEWLHIGSNLKLVFSFVLGLVSMLVLRP-