| DPOGS206754 | ||
|---|---|---|
| Transcript | DPOGS206754-TA | 642 bp |
| Protein | DPOGS206754-PA | 213 aa |
| Genomic position | DPSCF300316 + 14672-15392 | |
| RNAseq coverage | 1x (Rank: top 94%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL011205 | 1e-35 | 53.55% | |
| Bombyx | BGIBMGA009721-TA | 3e-15 | 35.06% | |
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_P43515 | 6e-16 | 40.53% | Chorion class B protein Ld10 n=4 Tax=Lymantria dispar RepID=CHB1_LYMDI |
| NCBI RefSeq | NP_001112374.1 | 4e-13 | 38.75% | chorion class CB protein M5H4 precursor [Bombyx mori] |
| NCBI nr blastp | gi|45476805 | 1e-15 | 40.53% | chorion protein [Lymantria dispar] |
| NCBI nr blastx | gi|116263 | 4e-26 | 43.16% | chorion protein [Antheraea polyphemus] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0007304 | 8e-26 | chorion-containing eggshell formation | |
| GO:0005213 | 8e-26 | structural constituent of chorion | ||
| GO:0007275 | 8e-26 | multicellular organismal development | ||
| GO:0042600 | 8e-26 | chorion | ||
| KEGG pathway | ||||
| InterPro domain | [3-182] IPR002635 | 8e-26 | Chorion protein | |
| Orthology group | MCL19879 | Lepidoptera specific | ||
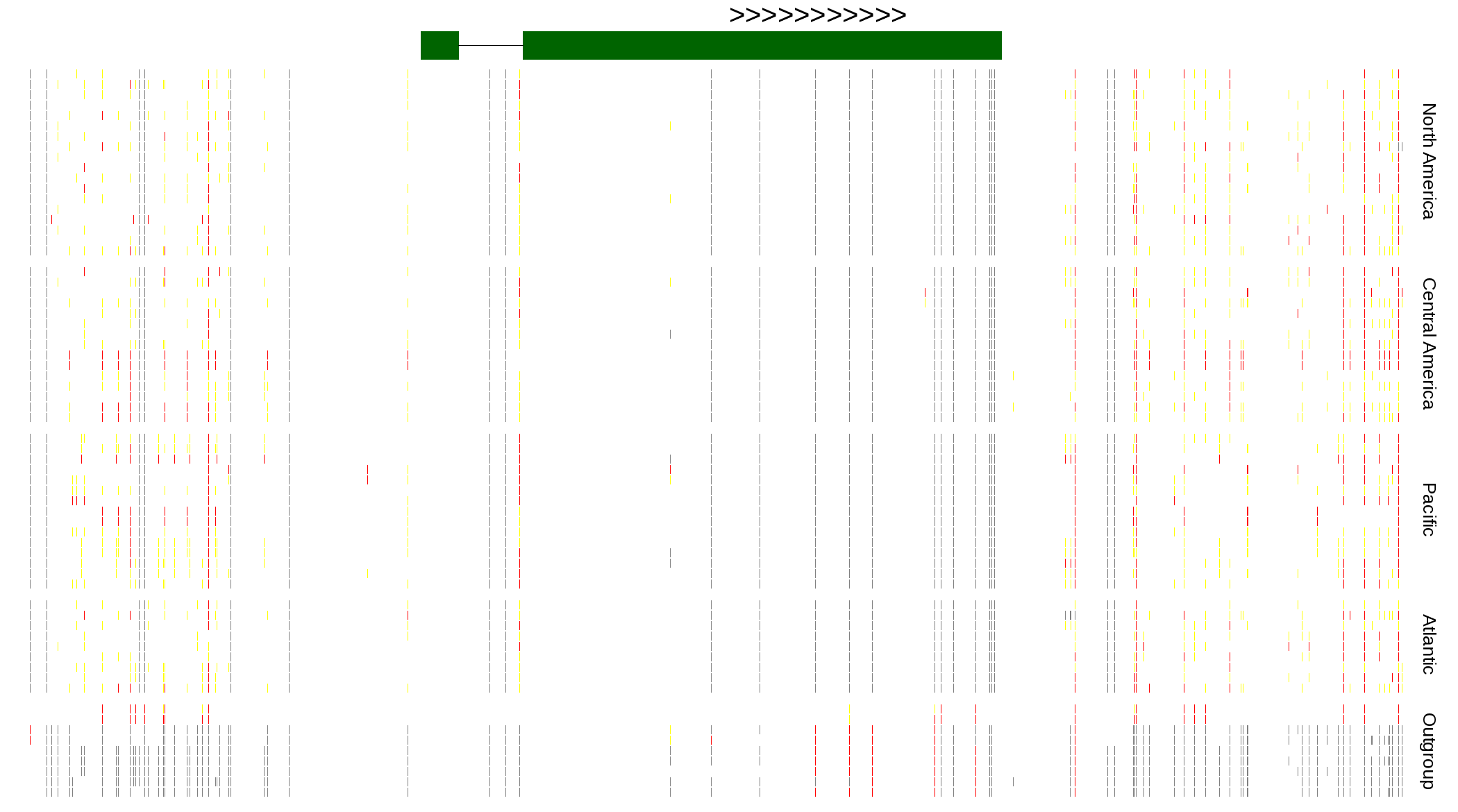
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206754-TA
ATGTTCAAGGCTACTCTCCTGGTATGCGCTCAGGCGCTCTTCGTTCAGTCCATCGCTGGACAGTGTCTTGGAGCTGGTTACGGTCCTCTCGCCGCTGAAATTCCTCTGTCTGCTGCCAACTGGGCTGGTATGTCTGCTGGTCCTTGCGGTGCTGCTGGTCTCTTTGACGGTTCTTGGTCTGCCGCCGGTGGTCCCTTCTATGGTGCTGGTTACGGACCTGCTGCCGCTTCTGCCTCTCATGGAGCTTTACCTGTATCCAGTGCATCCATGATCCCACCAAGTGGAGTGTCTGTACGATCAGACAACGCCATCGAAGGACCTCTCGCAGTCTCCGGTGCTCTGCCATTCCTGGGAACCGTAGCTTTGGAGGGCGCTCTGCCCACCGCTGGTGCTGGTGCAGTGGCTTACGGAGCCGGTAACGGAGAAGTCGCCATGTTGTCTGAAGATATCGGCGCCGATGGATTCAACTCCCTGGCTGGTGGTCTTGGTTATGGAGCTGGTGCTCTTGGTTATGGAGCTGGTGCTCTTGGCTATGGAGCTGGTGCCCTTGGCCTTGCAACTGGTCCTTTAGGATACAACGGTGCTTTATCTGGCCTAGGATATGCACGAACTGGTTGCGGCTGTGGATCCCGTCTCATCTAA
>DPOGS206754-PA
MFKATLLVCAQALFVQSIAGQCLGAGYGPLAAEIPLSAANWAGMSAGPCGAAGLFDGSWSAAGGPFYGAGYGPAAASASHGALPVSSASMIPPSGVSVRSDNAIEGPLAVSGALPFLGTVALEGALPTAGAGAVAYGAGNGEVAMLSEDIGADGFNSLAGGLGYGAGALGYGAGALGYGAGALGLATGPLGYNGALSGLGYARTGCGCGSRLI-