| DPOGS206773 | ||
|---|---|---|
| Transcript | DPOGS206773-TA | 564 bp |
| Protein | DPOGS206773-PA | 187 aa |
| Genomic position | DPSCF300001 - 6122247-6125294 | |
| RNAseq coverage | 9x (Rank: top 85%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006188 | 3e-54 | 56.61% | |
| Bombyx | BGIBMGA010844-TA | 1e-33 | 54.33% | |
| Drosophila | Or67c-PA | 1e-09 | 27.27% | |
| EBI UniRef50 | UniRef50_B7S6U6 | 7e-49 | 49.72% | Putative chemosensory receptor 12 n=3 Tax=Noctuidae RepID=B7S6U6_SPOLI |
| NCBI RefSeq | XP_321153.1 | 1e-12 | 29.14% | candidate olfactory receptor (AGAP001912-PA) [Anopheles gambiae str. PEST] |
| NCBI nr blastp | gi|148533561 | 2e-48 | 49.72% | putative chemosensory receptor 12 [Spodoptera littoralis] |
| NCBI nr blastx | gi|148533561 | 2e-47 | 49.72% | putative chemosensory receptor 12 [Spodoptera littoralis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0016020 | 3.1e-26 | membrane | |
| GO:0007608 | 3.1e-26 | sensory perception of smell | ||
| GO:0005549 | 3.1e-26 | odorant binding | ||
| GO:0004984 | 3.1e-26 | olfactory receptor activity | ||
| KEGG pathway | ||||
| InterPro domain | [3-178] IPR004117 | 3.1e-26 | Olfactory receptor, Drosophila | |
| Orthology group | MCL25782 | Lepidoptera specific | ||
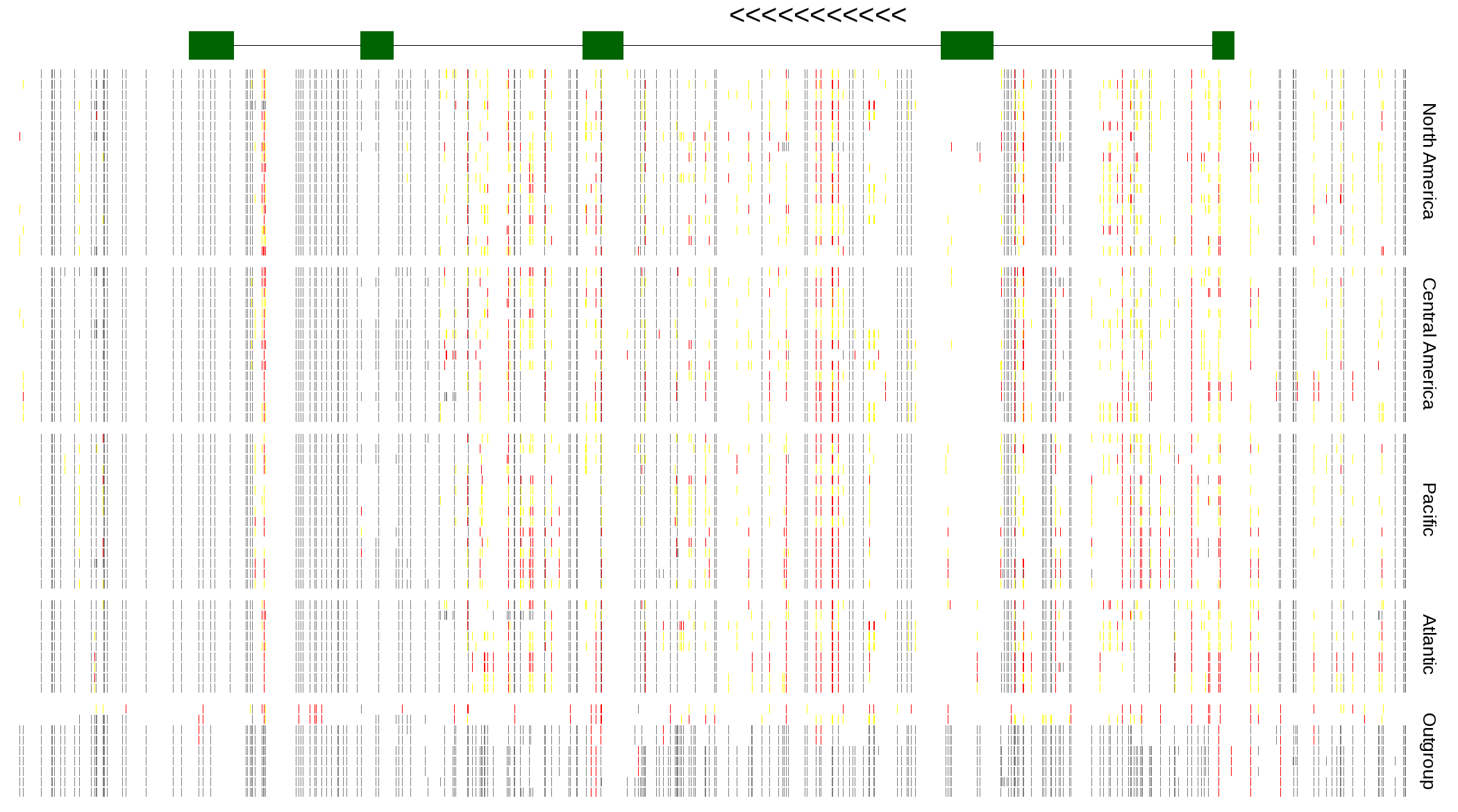
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS206773-TA
ATGCCATTCGATCCGTCTAAGAAGGTTGTCTTTGAAGTAGTACTTATTGTGCAAAACATACATTGTTTCCTCTCTGCAGCCTACATGTTAGCAGGAGATCTCTTGTTCTTCGTATTCCTCAGTCACATCACTACACAATTTTCCTTACTTGCTGTCAGGATTCAGAAGATGTTCTACGTTCCAATTGATGGACAGCTACCGGAGTCGTATCCACTCGGATATTATAACAAAGAAAATTCATTAAATAATCCAAATCAGGACACAGCAAACAACAGATGCAAGGTGGAAGAGGAGGAGCTTAAAAATATTGTCATACGACATAAAGCTTTAATTCGATTGTCAAATGACGTTGAGAACATGTATTCGTTTTCACTTCTTGTGAATTTCCTGAACAGCTCCATAATTATATGCTTCTGTTTATTTTGCTGCGCGTTCGTTGAAAAATGGAACGAGTTTAATTACAAAATATTTTTCATAACCGCCATTTCTCAGATATGGATACTCTGTTGGTACGGACAGCACTTGGTTGATACTGTTAGTTTATTTTATATTTCACTTGGTTAA
>DPOGS206773-PA
MPFDPSKKVVFEVVLIVQNIHCFLSAAYMLAGDLLFFVFLSHITTQFSLLAVRIQKMFYVPIDGQLPESYPLGYYNKENSLNNPNQDTANNRCKVEEEELKNIVIRHKALIRLSNDVENMYSFSLLVNFLNSSIIICFCLFCCAFVEKWNEFNYKIFFITAISQIWILCWYGQHLVDTVSLFYISLG-