| DPOGS207816 | ||
|---|---|---|
| Transcript | DPOGS207816-TA | 558 bp |
| Protein | DPOGS207816-PA | 185 aa |
| Genomic position | DPSCF300042 + 635422-636135 | |
| RNAseq coverage | 32x (Rank: top 75%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL011972 | 1e-66 | 61.96% | |
| Bombyx | % | |||
| Drosophila | agt-PA | 7e-11 | 39.77% | |
| EBI UniRef50 | UniRef50_D1AFU1 | 9e-26 | 36.53% | Methylated-DNA/protein-cysteine methyltransferase n=2 Tax=Bacteria RepID=D1AFU1_SEBTE |
| NCBI RefSeq | XP_003087247.1 | 1e-17 | 30.30% | hypothetical protein CRE_10797 [Caenorhabditis remanei] |
| NCBI nr blastp | gi|189501736 | 3e-34 | 39.62% | hypothetical protein Aasi_0286 [Candidatus Amoebophilus asiaticus 5a2] |
| NCBI nr blastx | gi|189501736 | 5e-32 | 39.62% | hypothetical protein Aasi_0286 [Candidatus Amoebophilus asiaticus 5a2] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0006281 | 8.4e-26 | DNA repair | |
| GO:0003824 | 8.4e-26 | catalytic activity | ||
| KEGG pathway | ||||
| InterPro domain | [100-179] IPR014048 | 8.4e-26 | Methylated-DNA-[protein]-cysteine S-methyltransferase, DNA binding | |
| [99-180] IPR011991 | 9.6e-26 | Winged helix-turn-helix transcription repressor DNA-binding | ||
| Orthology group | MCL22643 | Patchy | ||
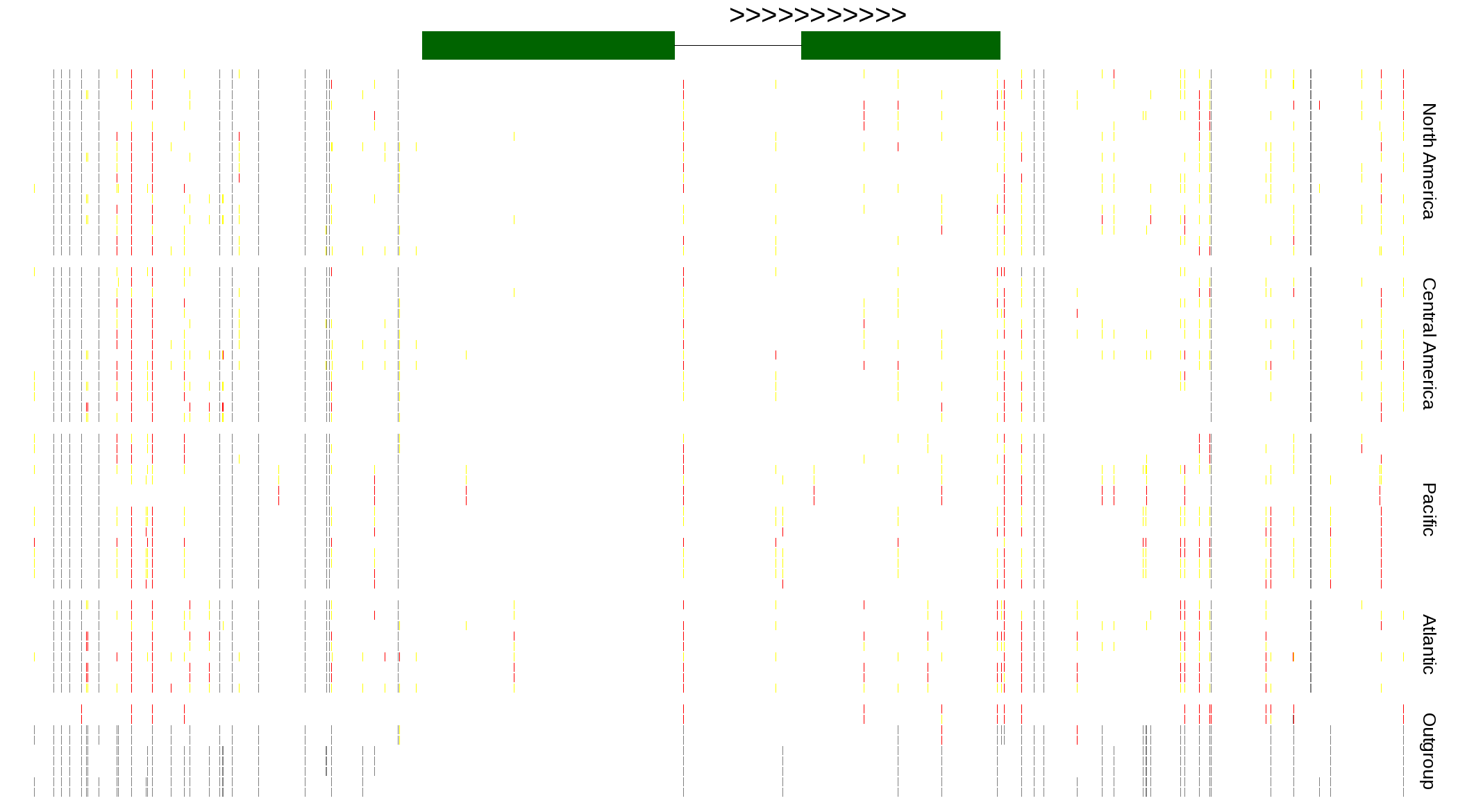
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS207816-TA
ATGGAGTTTTCAGAATTATTAACAAAGTTTTCTCAAAACTCTAAGAAATCCATTTGCATATTGAGATTTGATTCCCCTATTGGAAAAATAGTTGCAGCGGGTGATGATAACTTCCTATATGTTGTAGTTTTTGAAGACTCAAAAAACTTCGAGAAAGAATTACGTGTTATTGCTGATAATTTATCATGTAGTTTTGTAGAAAAAAAAAGTAAGATTCTAGAGCAGTTCGAGTGTGAAATCAAAGATTACTTCGATGGAAATTTAAGAAAGTTTACAGTTCCTATAAAAACTTTTGGTACTGATTTTCAAAAGGAAGTTTGGAAACACCTCCTGGAACTGCCCTATGGTTCTACACAGACCTATGGGTCACTTGCCAAGAATATGGGTAGGGCGGCAAGCCATTCTCGAGCTATTGGTGCAGCTTGTGGTGCGAATGCCCACCTTATAATAATACCTTGCCATCGTTTAGTTGCTTCTGCTTCTAACGGAGGATTTAGTTGTGGCGTTAATCGCAAGGAGTGGTTGATAGAGCATGAGAAGAAATTTGTTAAGCTTTGA
>DPOGS207816-PA
MEFSELLTKFSQNSKKSICILRFDSPIGKIVAAGDDNFLYVVVFEDSKNFEKELRVIADNLSCSFVEKKSKILEQFECEIKDYFDGNLRKFTVPIKTFGTDFQKEVWKHLLELPYGSTQTYGSLAKNMGRAASHSRAIGAACGANAHLIIIPCHRLVASASNGGFSCGVNRKEWLIEHEKKFVKL-