| DPOGS208353 | ||
|---|---|---|
| Transcript | DPOGS208353-TA | 651 bp |
| Protein | DPOGS208353-PA | 216 aa |
| Genomic position | DPSCF300251 - 49482-97769 | |
| RNAseq coverage | 102x (Rank: top 61%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL008044 | 7e-46 | 56.96% | |
| Bombyx | BGIBMGA014552-TA | 1e-19 | 64.29% | |
| Drosophila | nrm-PE | 2e-26 | 32.20% | |
| EBI UniRef50 | UniRef50_D7EJY4 | 6e-33 | 37.62% | Putative uncharacterized protein n=2 Tax=Tribolium castaneum RepID=D7EJY4_TRICA |
| NCBI RefSeq | XP_971833.2 | 6e-33 | 37.56% | PREDICTED: similar to AGAP012343-PA [Tribolium castaneum] |
| NCBI nr blastp | gi|270016467 | 2e-32 | 37.62% | hypothetical protein TcasGA2_TC006983 [Tribolium castaneum] |
| NCBI nr blastx | gi|270016467 | 2e-32 | 37.62% | hypothetical protein TcasGA2_TC006983 [Tribolium castaneum] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ptr:469866 | 9e-08 | ||
| K06548 (SN, SIGLEC1) | maps-> | Cell adhesion molecules (CAMs) | ||
| InterPro domain | [74-160] IPR013783 | 4.5e-09 | Immunoglobulin-like fold | |
| [81-139] IPR013151 | 2.5e-07 | Immunoglobulin | ||
| Orthology group | MCL26207 | Lepidoptera specific | ||
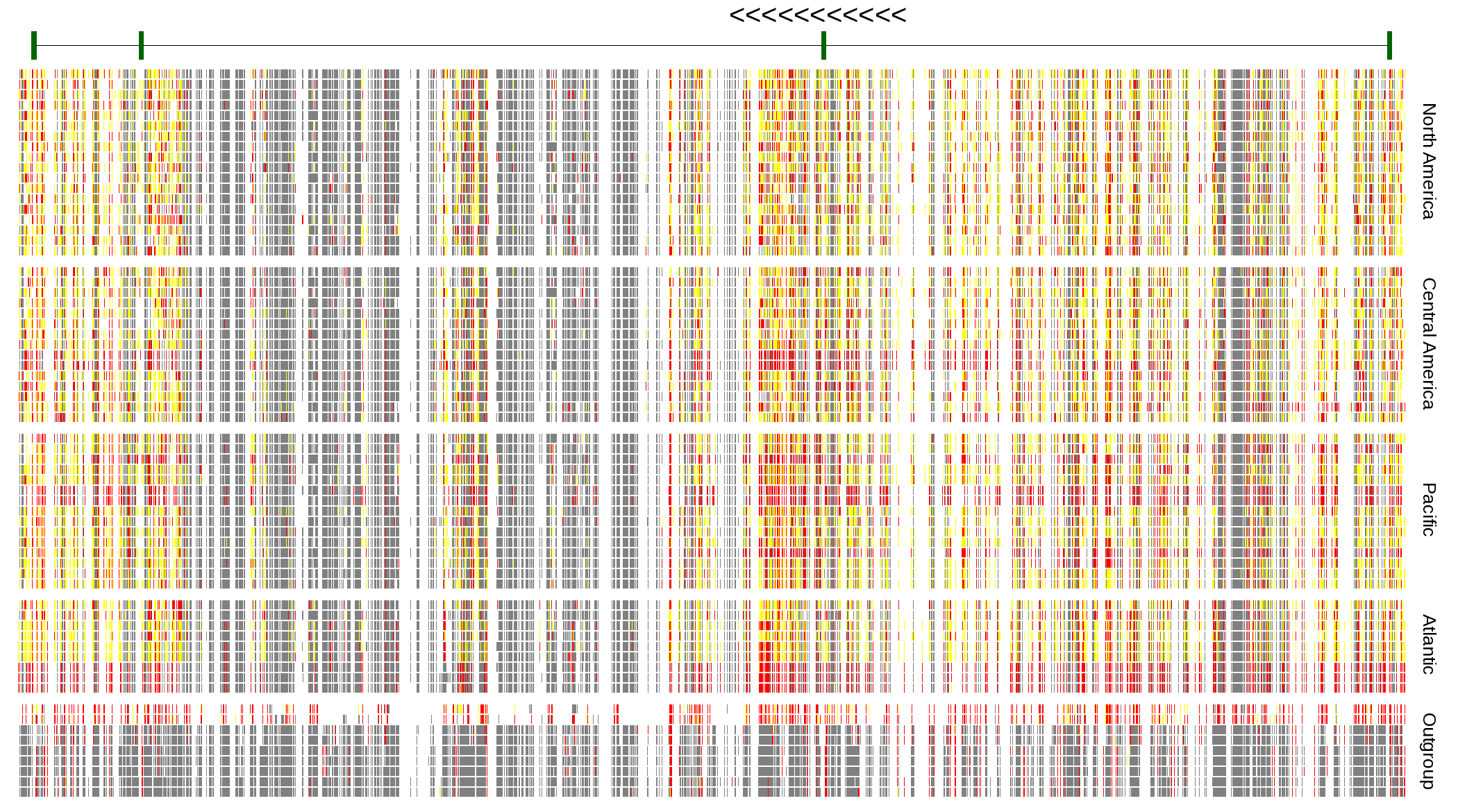
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS208353-TA
ATGTCGATATCCCACGAGCCTAATGCCACGTTCGCGGAACTGGTGATCACCAATGTTCATCCTGATGATGAAGGCGTATACCGCTGCGAAATCACCTACCTCCAAGTCGGCGAAGATTGCAACACGGTGCAAGTCACGGACTTCCATACATACATTAAACCGAAATCTGTGGAAGTGAGGACTGGAGAGGGTCTTATTTTGTCCGAGGGCGCCGTGTTTGGACCACTTCTAGAGAGGACGAAGGTTAATATTACGTGCCTGGTTGTAGGTGGCAAGCCTCAACCAAAAGTCACTTGGTATTTCAATGGAAAAGAAAGACTGGATGCCCAAACCATCACCGATCAATCATCGATAAAACATGTACTACCATTGACTGTGACACGAGCGGAGTTAGCTGGTGAACTCACTTGTAATGTGACCTCTGCTGCTATGGACAAGGCTGTGTTGAGAAAGGTTTTGTTGGATGTCAGAGTGCCGCCAAATAAGATCTCTATATCTGGAGTAGGCGAGCATGCTACCCAAGGTACGCTTCTGACGTTGATGTGTACGGTGTCCGGAGCTCGACCAGCGGCGAGGATTGATTGGTTTAATGGCACACACCAGCTTAGTCCAAAACAACAAAATATTCAAGTAAGTATAAAAAAGGAGTAG
>DPOGS208353-PA
MSISHEPNATFAELVITNVHPDDEGVYRCEITYLQVGEDCNTVQVTDFHTYIKPKSVEVRTGEGLILSEGAVFGPLLERTKVNITCLVVGGKPQPKVTWYFNGKERLDAQTITDQSSIKHVLPLTVTRAELAGELTCNVTSAAMDKAVLRKVLLDVRVPPNKISISGVGEHATQGTLLTLMCTVSGARPAARIDWFNGTHQLSPKQQNIQVSIKKE-