| DPOGS209497 | ||
|---|---|---|
| Transcript | DPOGS209497-TA | 300 bp |
| Protein | DPOGS209497-PA | 99 aa |
| Genomic position | DPSCF300127 - 165686-167158 | |
| RNAseq coverage | 0x (Rank: top 98%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL016024 | 6e-20 | 53.42% | |
| Bombyx | BGIBMGA007458-TA | 1e-08 | 41.30% | |
| Drosophila | CG6421-PA | 1e-07 | 37.14% | |
| EBI UniRef50 | UniRef50_B6RQP9 | 1e-11 | 47.89% | I-type lysozyme n=2 Tax=Cucujiformia RepID=B6RQP9_SITZE |
| NCBI RefSeq | XP_973861.2 | 8e-11 | 41.56% | PREDICTED: similar to CG6426 CG6426-PA [Tribolium castaneum] |
| NCBI nr blastp | gi|167444218 | 4e-11 | 47.89% | i-type lysozyme [Sitophilus zeamais] |
| NCBI nr blastx | gi|167444218 | 4e-12 | 47.89% | i-type lysozyme [Sitophilus zeamais] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003796 | 2.4e-12 | lysozyme activity | |
| KEGG pathway | ||||
| InterPro domain | [26-71] IPR008597 | 2.4e-12 | Destabilase | |
| Orthology group | ||||
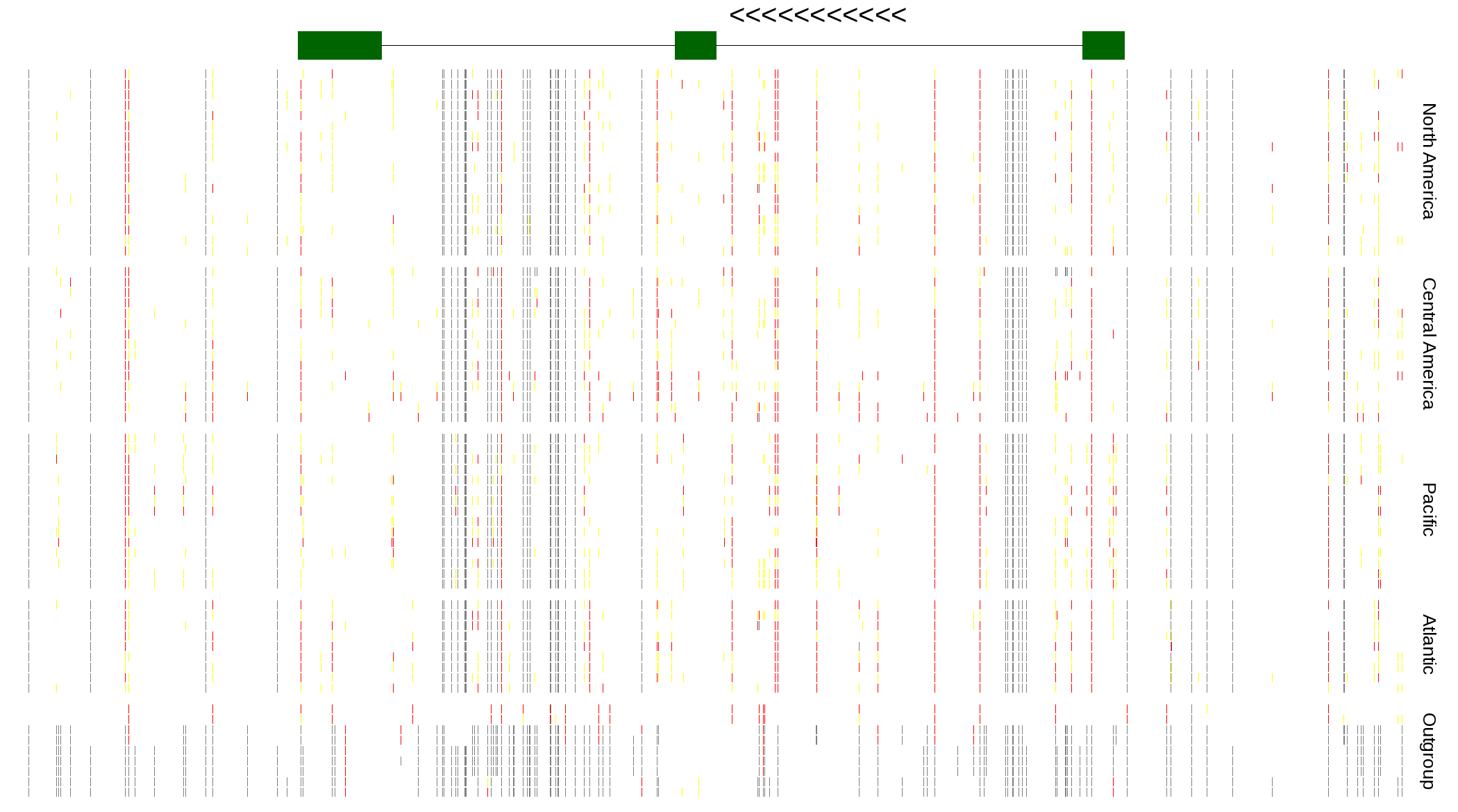
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS209497-TA
ATGTTTACCGACACAAAGACATCAATACTATCACTTCTGTTGATCACAGCTAGTTTGTTGTTTATTGAATCTGCAGCTTGGAAGGATTGCGCTAATGACTACTTCTGCGCTCAGGACATAATGAAAGGCTATCTTCAAAAGTTCTCAAAGGACTGCAATAGTGATGGCATGATCAATTGCTATGACATCTTGACGATTAATAGTAACGGCGGAGACTGCAGGCCGCTAAATCAGTCTTCGAGTGGACGCGTGTGGCTCAAACGTTACGAGGAATGTCGTGTAGCCAGAATTCTGACTTGA
>DPOGS209497-PA
MFTDTKTSILSLLLITASLLFIESAAWKDCANDYFCAQDIMKGYLQKFSKDCNSDGMINCYDILTINSNGGDCRPLNQSSSGRVWLKRYEECRVARILT-