| DPOGS209622 | ||
|---|---|---|
| Transcript | DPOGS209622-TA | 684 bp |
| Protein | DPOGS209622-PA | 227 aa |
| Genomic position | DPSCF300015 + 637524-638653 | |
| RNAseq coverage | 451x (Rank: top 27%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL017015 | 3e-106 | 80.09% | |
| Bombyx | BGIBMGA006687-TA | 1e-90 | 71.00% | |
| Drosophila | CG12393-PC | 6e-42 | 52.41% | |
| EBI UniRef50 | UniRef50_E2A9V1 | 2e-64 | 56.14% | Protein FAM109A n=10 Tax=Neoptera RepID=E2A9V1_CAMFO |
| NCBI RefSeq | XP_395804.2 | 3e-67 | 56.96% | PREDICTED: similar to CG12393-PA, isoform A [Apis mellifera] |
| NCBI nr blastp | gi|322802120 | 4e-67 | 58.44% | hypothetical protein SINV_06092 [Solenopsis invicta] |
| NCBI nr blastx | gi|156546916 | 4e-64 | 57.69% | PREDICTED: sesquipedalian-1-like [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005515 | 7.4e-23 | protein binding | |
| KEGG pathway | nvi:100116145 | 1e-10 | ||
| K01109 (INPP4) | maps-> | Phosphatidylinositol signaling system | ||
| Inositol phosphate metabolism | ||||
| InterPro domain | [17-111] IPR011993 | 7.4e-23 | Pleckstrin homology-type | |
| [18-114] IPR001849 | 2.8e-19 | Pleckstrin homology domain | ||
| Orthology group | MCL12478 | Single-copy universal gene | ||
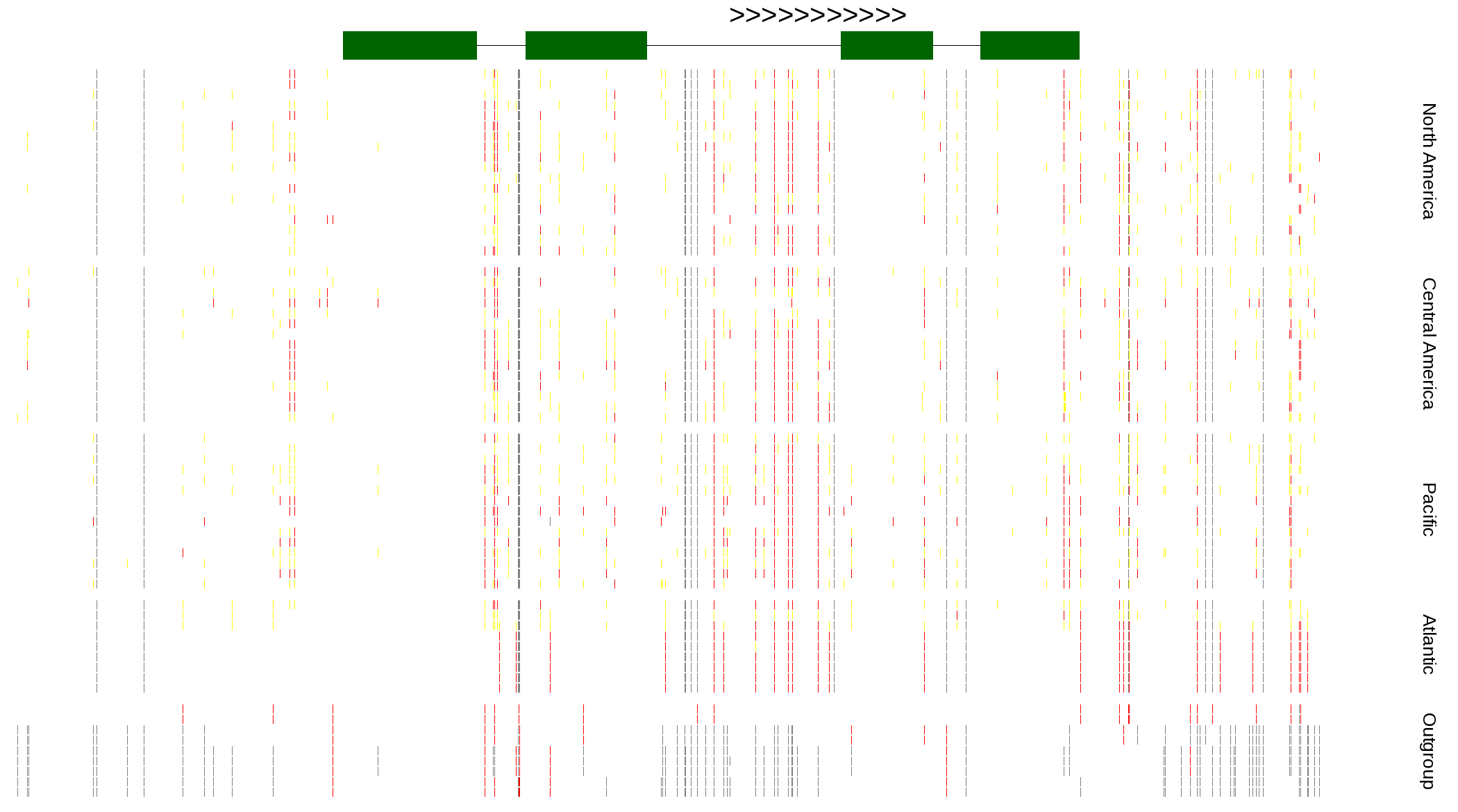
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS209622-TA
ATGAAGATTAACGAGAAAAATTTATGTGCTTTCGCTTCATCTTCAACCCCCGTGGATCGGGAGGGTTGGTTAGAAATGAGGGGTGAAGTCGGTAAGAGTTACCAAAGAAGATTTTTTATGTTAAAAGGGAATCTGCTGTTTTACTTTGAAAAGAAAGGAGACAAAGAACCCGTAGGGGTCGTTATACTCGAGGGATGCACAATTGAATTAACAGAGGAAGAATCATATAGCTTCAAAATAGTATTCCACTGTGAAGGTGGCCGTACTTATTACTTATGCACAAACTCACAGTCGTCAATGGAGGGTTGGATGAAAGCCTTGGCCTGTGCAAGCTATGATTATATGAAACTGATGGTAGCTGAGCTTCAGAGGCAGTTGGATGAAGCTCAAGCTGAAGAATCTGTAGAAGCTGCGACTGTGGTCACCGTCTCTCAAGCAGGGGAAGAACCCAAAGTACCACCGAGAGGACAGAGACACAACCCATTTAACAAAGCACCTGATGATAGTAAAGTTGTTACCGGGAGTAGACATTACAAAGAGAACATGAGGCCAGAAGTGGCTCGAAAGAAAGTTCCATTCAGGGAAATACACAAAGCACACGGACGAAAGATCTTGTTGGACCGCAGTGAGTGGCGAGCCTCACTACGTGCGAGACTCAGTGAAACACCACTCATACAACTATGA
>DPOGS209622-PA
MKINEKNLCAFASSSTPVDREGWLEMRGEVGKSYQRRFFMLKGNLLFYFEKKGDKEPVGVVILEGCTIELTEEESYSFKIVFHCEGGRTYYLCTNSQSSMEGWMKALACASYDYMKLMVAELQRQLDEAQAEESVEAATVVTVSQAGEEPKVPPRGQRHNPFNKAPDDSKVVTGSRHYKENMRPEVARKKVPFREIHKAHGRKILLDRSEWRASLRARLSETPLIQL-