| DPOGS210076 | ||
|---|---|---|
| Transcript | DPOGS210076-TA | 627 bp |
| Protein | DPOGS210076-PA | 208 aa |
| Genomic position | DPSCF300017 - 149121-154740 | |
| RNAseq coverage | 133x (Rank: top 56%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006529 | 2e-12 | 52.17% | |
| Bombyx | BGIBMGA012742-TA | 5e-71 | 68.39% | |
| Drosophila | vn-PA | 2e-17 | 27.07% | |
| EBI UniRef50 | UniRef50_UPI0002062356 | 2e-29 | 43.40% | UPI0002062356 related cluster n=2 Tax=unknown RepID=UPI0002062356 |
| NCBI RefSeq | XP_002027988.1 | 9e-22 | 32.39% | GL21406 [Drosophila persimilis] |
| NCBI nr blastp | gi|328786644 | 9e-33 | 47.88% | PREDICTED: hypothetical protein LOC408297 [Apis mellifera] |
| NCBI nr blastx | gi|383858253 | 4e-33 | 48.19% | PREDICTED: uncharacterized protein LOC100881392 [Megachile rotundata] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005515 | 7.2e-05 | protein binding | |
| KEGG pathway | tgu:100219062 | 5e-15 | ||
| K05456 (NRG2) | maps-> | ErbB signaling pathway | ||
| InterPro domain | [34-130] IPR013783 | 2.8e-21 | Immunoglobulin-like fold | |
| [39-116] IPR013098 | 7.9e-16 | Immunoglobulin I-set | ||
| [47-116] IPR003598 | 2.9e-14 | Immunoglobulin subtype 2 | ||
| [41-128] IPR003599 | 2.2e-11 | Immunoglobulin subtype | ||
| Orthology group | MCL18390 | Insect specific | ||
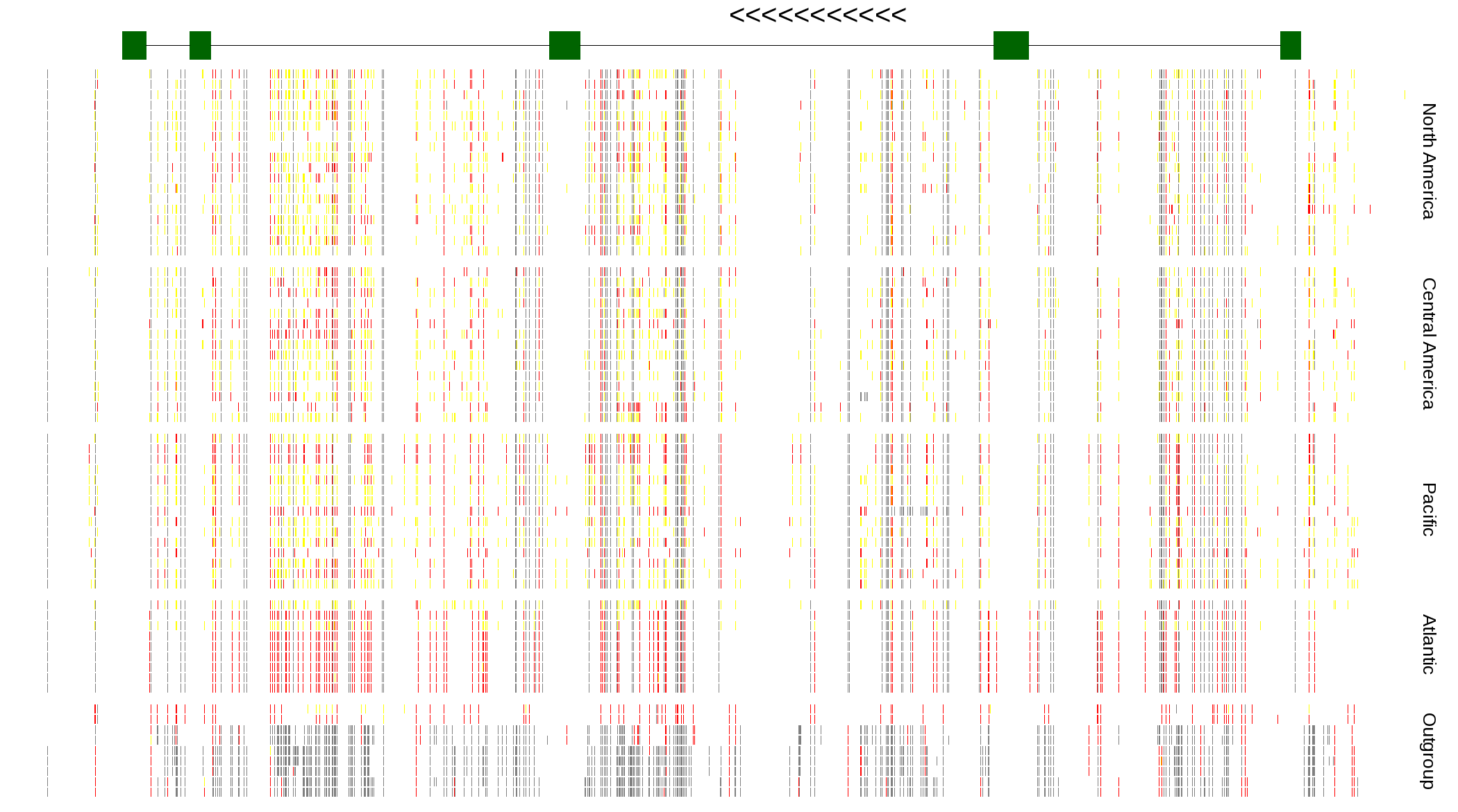
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS210076-TA
ATGGATGGTTCATTCATATTAAAGTTAGTACGTCAGTCTGTAGGCGAGTCCATTGGCAAGGCGAGGAATTCCGTTTTAGAGCGTATTATGAATGAAAGTAGCCCCGCTCGGATAGAGCCAATGCAATCTGTTCGTATTTCTCGAGGAAGTAGAGTGAAATTGGAGTGTCGAGCCAGTGCTCGTCCTCCGCCGAGGATATCATGGTACAGGGATGGCCGGCCAGTCGCCGAAATCGCACTCAGGCGGTTCCGGGTGCAAAATTATAGACGTCGCTCGATCCTCGTGATCCGCCACGCTCGTCGCGAGGACTCCGCCACATACGAGTGTCGAGCCCAGGGCGCTATAGGACCTCCTGCTGTAACCAGCGCTAATGTCAGCGTCGTGGCACCAGTCGCCACCGCACCGGATACTTCGATCCTCGGCGCCCCTTGTCCGATGCCTGACCCAGCCTCCTACTGTCTGAATGGAGGCACTTGTTTATTCTTTGAGCTCGTTCAAGAGCAAGCTTGCAAATGCCCGGAAGGCTTCAACGGGCAACGCTGCGAAAATAAGGACGTGTCAAATAGGAGCAGTATGTATCATTCCTACACATGCAAACTGGGCCTCTCCACGTCCTACTACTGCTAG
>DPOGS210076-PA
MDGSFILKLVRQSVGESIGKARNSVLERIMNESSPARIEPMQSVRISRGSRVKLECRASARPPPRISWYRDGRPVAEIALRRFRVQNYRRRSILVIRHARREDSATYECRAQGAIGPPAVTSANVSVVAPVATAPDTSILGAPCPMPDPASYCLNGGTCLFFELVQEQACKCPEGFNGQRCENKDVSNRSSMYHSYTCKLGLSTSYYC-