| DPOGS210540 | ||
|---|---|---|
| Transcript | DPOGS210540-TA | 654 bp |
| Protein | DPOGS210540-PA | 217 aa |
| Genomic position | DPSCF300304 - 149632-151496 | |
| RNAseq coverage | 192x (Rank: top 48%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL009547 | 4e-129 | 99.08% | |
| Bombyx | BGIBMGA013463-TA | 7e-128 | 98.16% | |
| Drosophila | mats-PA | 2e-119 | 92.17% | |
| EBI UniRef50 | UniRef50_Q7L9L4 | 9e-115 | 89.30% | MOB kinase activator 1B n=227 Tax=root RepID=MOB1B_HUMAN |
| NCBI RefSeq | XP_393046.2 | 7e-124 | 95.85% | PREDICTED: similar to mob as tumor suppressor CG13852-PA [Apis mellifera] |
| NCBI nr blastp | gi|345489125 | 1e-122 | 95.85% | PREDICTED: mps one binder kinase activator-like 1-like [Nasonia vitripennis] |
| NCBI nr blastx | gi|345489125 | 3e-120 | 95.85% | PREDICTED: mps one binder kinase activator-like 1-like [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ame:409540 | 2e-123 | ||
| K06685 (MOB1) | maps-> | Cell cycle - yeast | ||
| InterPro domain | [1-216] IPR005301 | 1.7e-161 | Mob1/phocein | |
| Orthology group | MCL12409 | Multiple-copy universal gene | ||
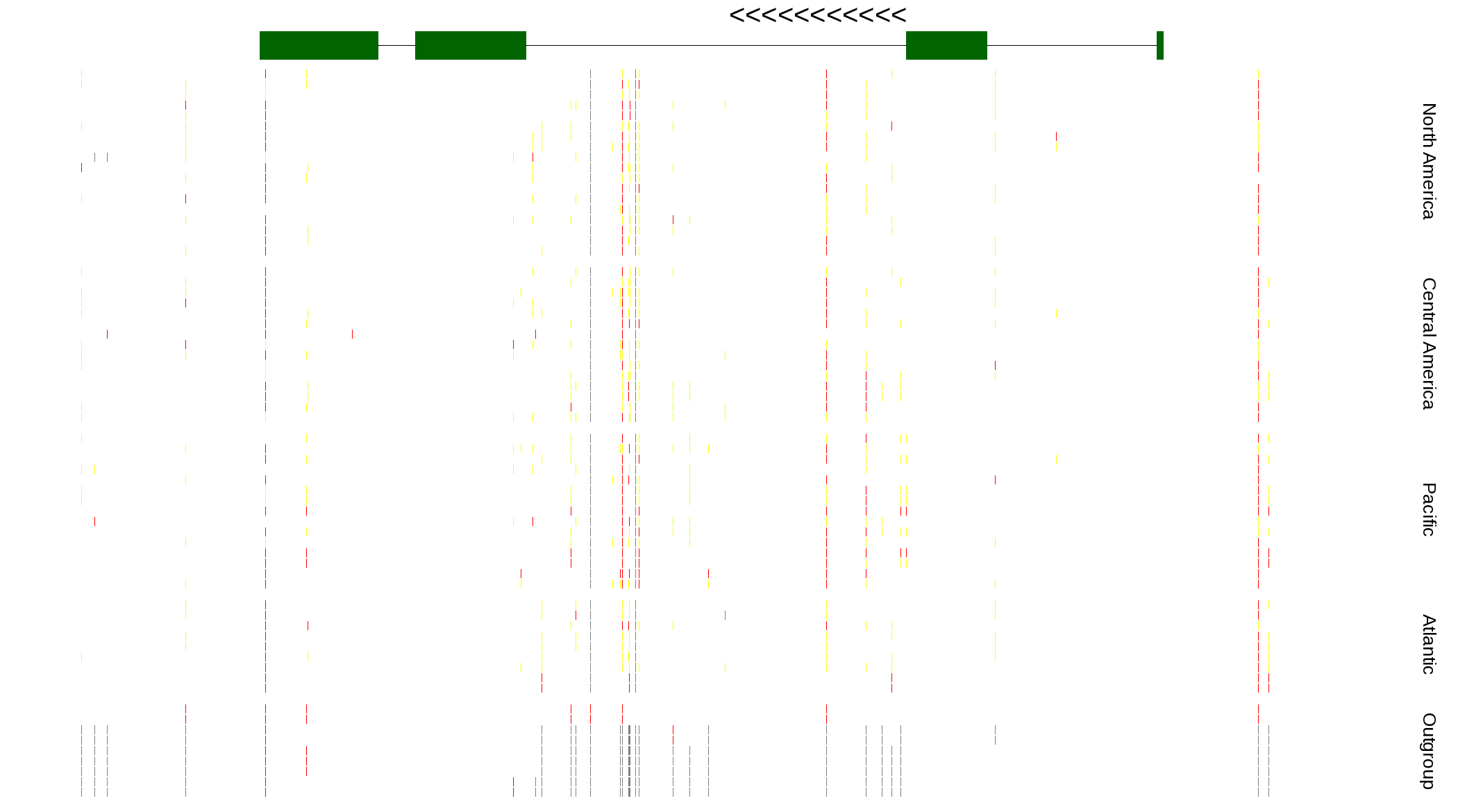
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS210540-TA
ATGAGCTTTCTATTTGGAAGTCGATCGAACAAAACCTTCAAGCCAAAGAAAAATATACCTGAGGGAACTCATCAGTATGACCTGATGCGTCACGCGGCGGCCACACTGGGTTCAGGTAACCTACGACTGGCTGTCATGCTGCCTGAGGGAGAGGATCTCAATGAATGGGTTGCAGTTAACACCGTAGACTTCTTCAACCAAATCAATATGCTGTATGGAACCATCACGGAGTTCTGCACGGAGGAGTCATGCGCCGTGATGTCCGCGGGACCCAAGTACGAGTACCACTGGGCCGACGGACACACCGTCAAGAAACCCATCAAGTGTTCCGCGCCCAAGTACATAGACTACCTCATGACGTGGGCGCAGGACCAGCTCGACGACGAGACGCTGTTCCCTTCCAAGATAGGAGTGCCGTTCCCCAAGAACTTCCTCTCGATGGCAAAAACGATCCTGAAGCGTCTGTTTCGCGTCTACGCCCACATCTACCACCAGCACTTCCCGGAGGTGGTGCAGCTGGGCGAGGAGGCGCACCTCAACACGTCCTTCAAGCACTTCATCTTCTTCGTGCAAGAGTTCAACCTCATCGAGCGCCGCGAGCTGGCCCCGCTGCAGGAGCTCATCGACAAACTCACCGCCAAAGAGGCGCGATGA
>DPOGS210540-PA
MSFLFGSRSNKTFKPKKNIPEGTHQYDLMRHAAATLGSGNLRLAVMLPEGEDLNEWVAVNTVDFFNQINMLYGTITEFCTEESCAVMSAGPKYEYHWADGHTVKKPIKCSAPKYIDYLMTWAQDQLDDETLFPSKIGVPFPKNFLSMAKTILKRLFRVYAHIYHQHFPEVVQLGEEAHLNTSFKHFIFFVQEFNLIERRELAPLQELIDKLTAKEAR-