| DPOGS210692 | ||
|---|---|---|
| Transcript | DPOGS210692-TA | 495 bp |
| Protein | DPOGS210692-PA | 164 aa |
| Genomic position | DPSCF300013 - 624068-626062 | |
| RNAseq coverage | 637x (Rank: top 20%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL020585 | 4e-39 | 64.00% | |
| Bombyx | BGIBMGA006308-TA | 3e-79 | 86.36% | |
| Drosophila | chp-PA | 1e-39 | 49.67% | |
| EBI UniRef50 | UniRef50_E2A0X8 | 4e-39 | 53.79% | Chaoptin n=8 Tax=Formicidae RepID=E2A0X8_CAMFO |
| NCBI RefSeq | XP_395331.3 | 6e-45 | 57.89% | PREDICTED: similar to Chaoptin precursor (Photoreceptor cell-specific membrane protein) [Apis mellifera] |
| NCBI nr blastp | gi|345481495 | 2e-44 | 55.84% | PREDICTED: chaoptin-like isoform 1 [Nasonia vitripennis] |
| NCBI nr blastx | gi|345481495 | 9e-44 | 55.84% | PREDICTED: chaoptin-like isoform 1 [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | ||||
| Orthology group | MCL34843 | Insect specific | ||
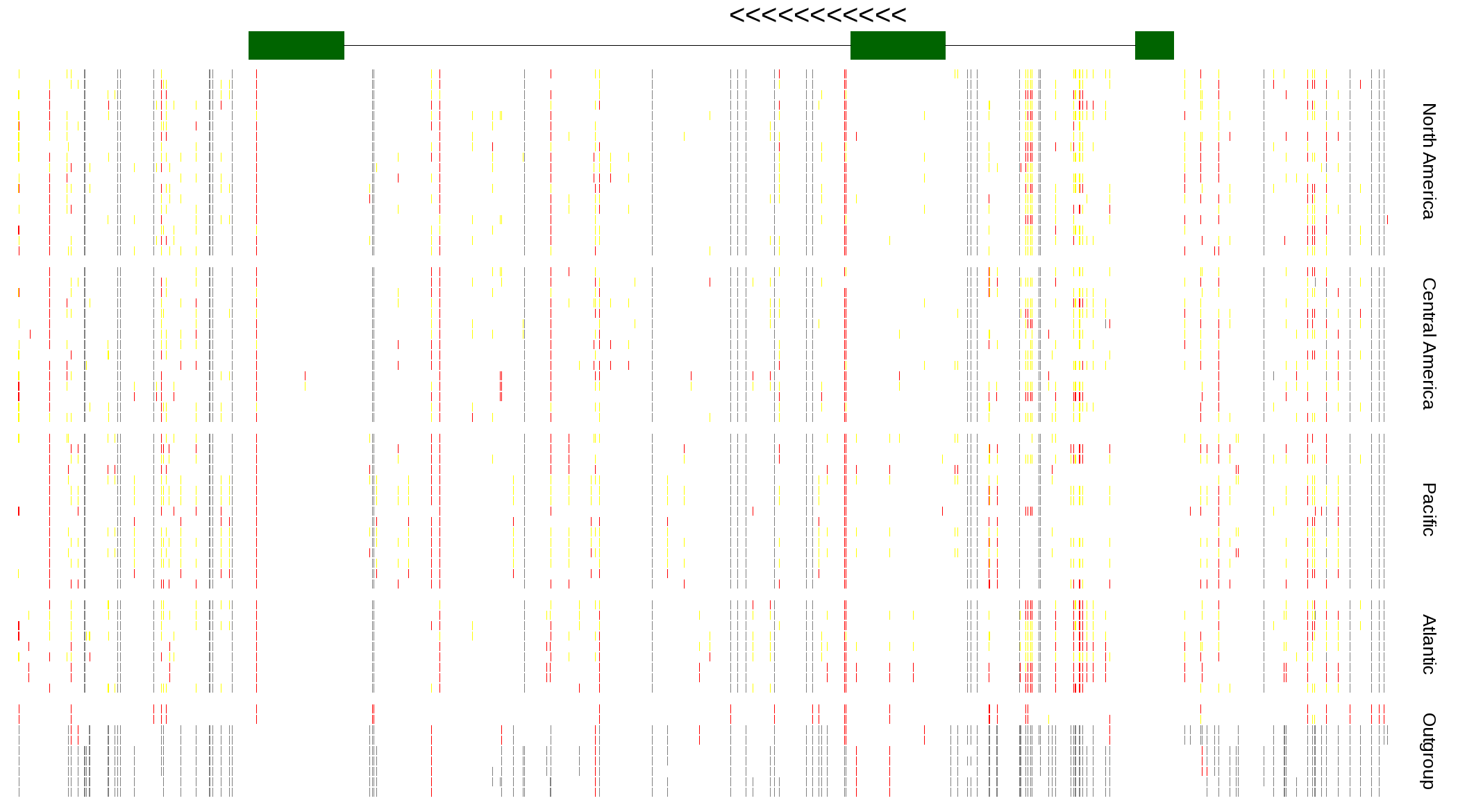
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS210692-TA
ATGACCTTAGTGAAATTCGGCTACACCCTTGTGGTAGTAACATTTTTGTTGATGATATGGGCGTCCTTAGCCAGGGCGTTAGAATTACACGTAGACACTGGACATCCGCCATGCCTTTTCCAAGCTCTCTGTACGTGCTCTAAACCTGATGGTGATCTTGGTATCGTGACGTGCAACCACGTGCCCATCCTCAGGGTACCAGCCGCCGTAAATAGCTCGAAAGTTTTCACGTTACAGTTGACTGGGAACAACATAAGAGATCTCGAACCGCAGTTCTTCCAAGCCACAGGGTTATACAGGCTGGCTATAAACCAGAATCCGCTGGAGAGTATACACGAGGAAGCTTTTTATGGCTTAGACAATACACTATGGGAGTTGGAATTAATGCAGGACAAGTTGACGGCTGTCCCGAGTCGAGCTCTGCGATATCTGCAAAAGCTGCGCCTCCTTGATTTGACGGGTGAGTGTCATAAAATTGTGGTTTTAAATGTCTAG
>DPOGS210692-PA
MTLVKFGYTLVVVTFLLMIWASLARALELHVDTGHPPCLFQALCTCSKPDGDLGIVTCNHVPILRVPAAVNSSKVFTLQLTGNNIRDLEPQFFQATGLYRLAINQNPLESIHEEAFYGLDNTLWELELMQDKLTAVPSRALRYLQKLRLLDLTGECHKIVVLNV-