| DPOGS211441 | ||
|---|---|---|
| Transcript | DPOGS211441-TA | 531 bp |
| Protein | DPOGS211441-PA | 176 aa |
| Genomic position | DPSCF300223 - 120184-120937 | |
| RNAseq coverage | 912x (Rank: top 14%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL007456 | 4e-68 | 69.01% | |
| Bombyx | BGIBMGA002190-TA | 2e-32 | 55.75% | |
| Drosophila | CG4210-PA | 1e-31 | 42.68% | |
| EBI UniRef50 | UniRef50_D6WL85 | 2e-36 | 47.83% | Putative uncharacterized protein n=1 Tax=Tribolium castaneum RepID=D6WL85_TRICA |
| NCBI RefSeq | XP_002053370.1 | 3e-37 | 50.00% | GJ23842 [Drosophila virilis] |
| NCBI nr blastp | gi|383860951 | 6e-37 | 48.77% | PREDICTED: diamine acetyltransferase 2-like [Megachile rotundata] |
| NCBI nr blastx | gi|383860951 | 1e-34 | 48.77% | PREDICTED: diamine acetyltransferase 2-like [Megachile rotundata] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0008152 | 3.6e-10 | metabolic process | |
| GO:0008080 | 3.6e-10 | N-acetyltransferase activity | ||
| KEGG pathway | tca:662606 | 1e-36 | ||
| K00657 (E2.3.1.57, speG) | maps-> | Arginine and proline metabolism | ||
| InterPro domain | [12-144] IPR016181 | 1e-28 | Acyl-CoA N-acyltransferase | |
| [74-143] IPR000182 | 3.6e-10 | GCN5-related N-acetyltransferase (GNAT) domain | ||
| Orthology group | MCL11970 | Single-copy universal gene | ||
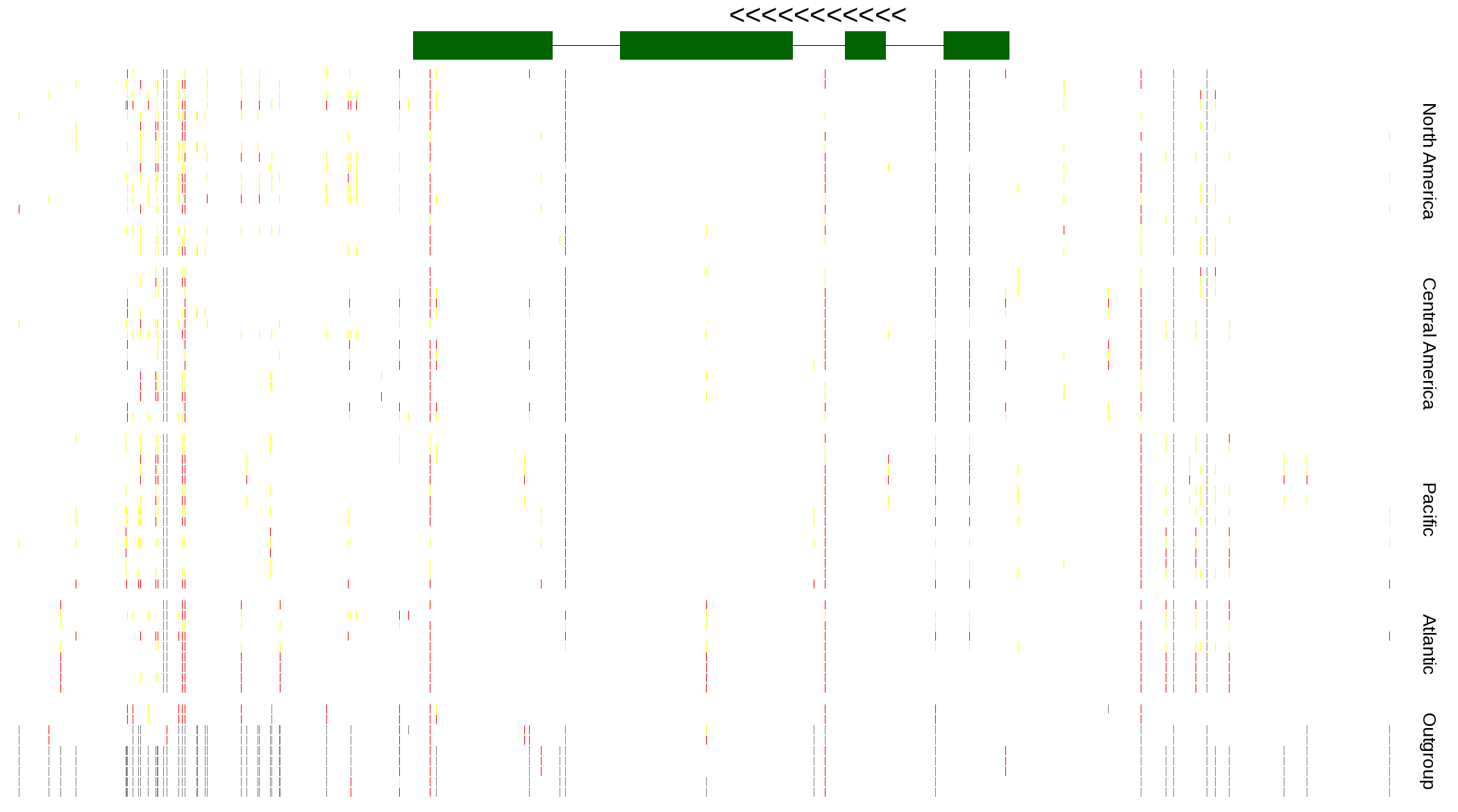
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS211441-TA
ATGGCCGCCAAATATGAGGACGGAGAAGTGAAGGTGCGGCGGTCCACGAGAGACGATATGAAGGCTGTCGCTGAAATGATACAGGAACTCGCGGATTTGGAGGAAATGTCGGACGGACCGAAACTGAGTGTGCAGGACTTGCAACGAGACGGCTTCGATCTGCAGCCGCCAGCGTTCTTGTGTCTGGTTGCGGAAGTTAACAAGGACGGTGCTCCTCTAGTAGCCGGCTATGCTTTATATTTCCCGGTTTATTCAACCTGGAATGGTCAGGCCATGATGTTAGAGGATCTGTATGTGCGGGCCAGTGAGCGGCGTCGGGGGCTCGGACATAGATTGTTTAATGCAGTCGCTAGAGAAGCTCACTTGATGGGTTCATCTAGACTAGACTTTCATGTTCTGGAGTGGAATAAAAATGCTAGAGATTTCTATGACTTGAAAGGAGCCATCAACATAACACAAACTGAGAAGTGGTGCTTCTACCGTCTCAGCGGTGATGCTTTGAAGAAGGTTGCTCTCGCTGCTAAATCATGA
>DPOGS211441-PA
MAAKYEDGEVKVRRSTRDDMKAVAEMIQELADLEEMSDGPKLSVQDLQRDGFDLQPPAFLCLVAEVNKDGAPLVAGYALYFPVYSTWNGQAMMLEDLYVRASERRRGLGHRLFNAVAREAHLMGSSRLDFHVLEWNKNARDFYDLKGAINITQTEKWCFYRLSGDALKKVALAAKS-