| DPOGS212432 | ||
|---|---|---|
| Transcript | DPOGS212432-TA | 624 bp |
| Protein | DPOGS212432-PA | 207 aa |
| Genomic position | DPSCF300258 + 109728-114352 | |
| RNAseq coverage | 78x (Rank: top 65%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL012357 | 2e-91 | 73.87% | |
| Bombyx | BGIBMGA002890-TA | 4e-62 | 62.18% | |
| Drosophila | GLaz-PA | 8e-42 | 40.31% | |
| EBI UniRef50 | UniRef50_A1Z991 | 9e-40 | 40.31% | Glial lazarillo n=15 Tax=Diptera RepID=A1Z991_DROME |
| NCBI RefSeq | XP_001661562.1 | 1e-47 | 44.33% | apolipoprotein D, putative [Aedes aegypti] |
| NCBI nr blastp | gi|157128947 | 2e-46 | 44.33% | apolipoprotein D, putative [Aedes aegypti] |
| NCBI nr blastx | gi|157128947 | 2e-44 | 44.72% | apolipoprotein D, putative [Aedes aegypti] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005488 | 6.8e-29 | binding | |
| GO:0006810 | 1.6e-07 | transport | ||
| GO:0005215 | 1.6e-07 | transporter activity | ||
| KEGG pathway | ||||
| InterPro domain | [29-200] IPR011038 | 1.6e-33 | Calycin-like | |
| [19-199] IPR012674 | 6.8e-29 | Calycin | ||
| [11-203] IPR022271 | 2.1e-09 | Lipocalin, ApoD type | ||
| [38-54] IPR003057 | 1.6e-07 | Invertebrate colouration protein | ||
| Orthology group | MCL18401 | Insect specific | ||
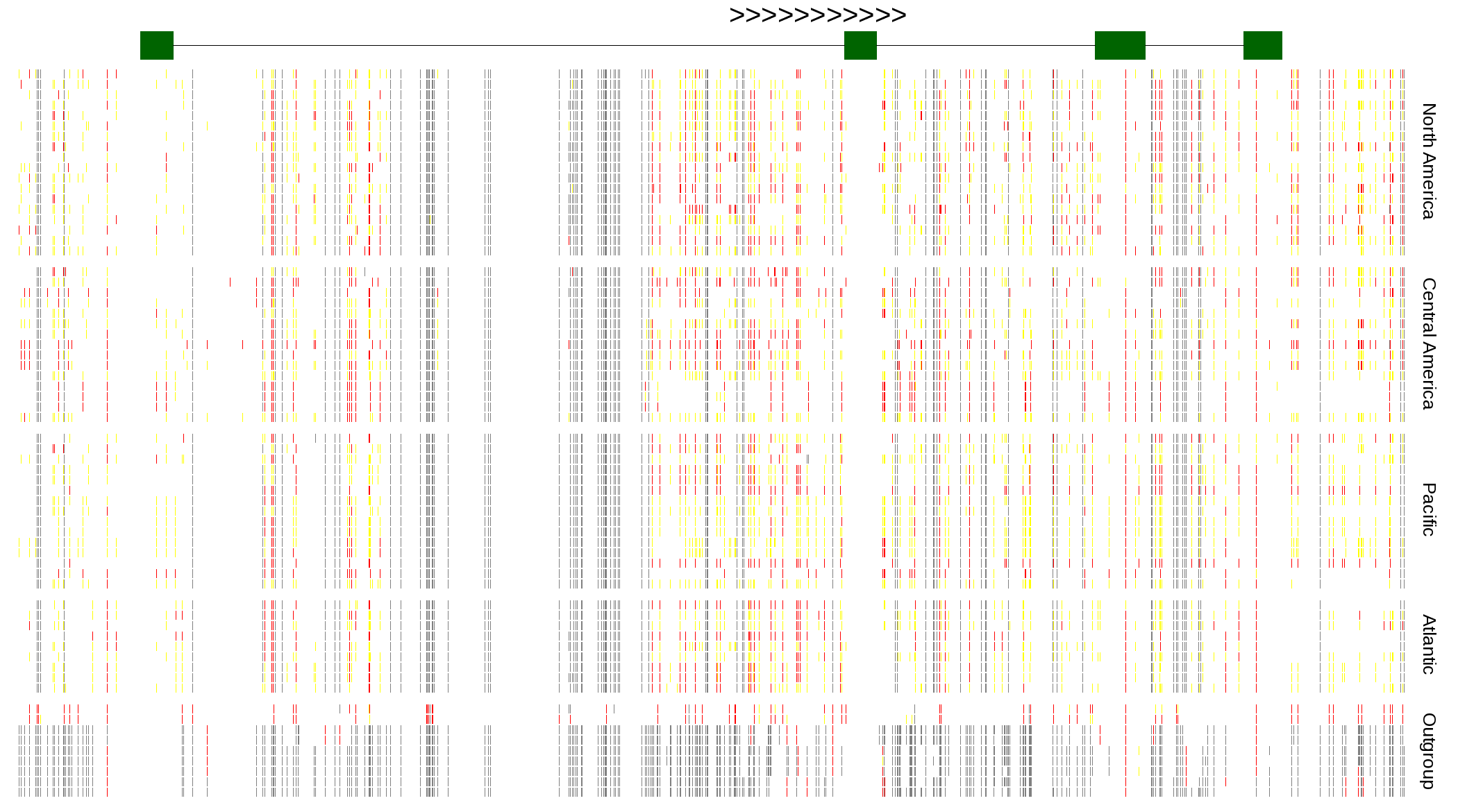
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS212432-TA
ATGGGTGACTTTCTAGTGATGGGAATCAAGTTGAAAATTATATTGTTGTGTACCGCAGTTAAAACTGTTACTAGCACCGGTTTAGGAAAATGCCCGATTTATCCACAGTTCCCAGATTATGACATCAATCGGATGACAGGGACCTGGTACGAGGTGGAGCGTTCCTTCCACTTGATGGAGATATCCGCAAGCTGCATTCAGCTGGACGTAACAATAAATGAAAGAGGATTTCTACTCATCACAGTCAACACTATCAATAGATGGACAAACACGCCGTCCATTAGCTACGGATTGGGTATTCCTAGTCATAACGGCTCGTCTTCCTTCCGCTACAAGTTGAACAATCGTATGCCGTACATCATCGGTCGCATGCTGCCCGGAGCCGGTCTCTACAACGTACTTTTTACTGACTACGAACGATTTGCAATAATTTGGTCCTGCACTAACTATAGTATAGCACATGCAGACCGTCTGTGGATCTTAGGCCGTCACCGGTCTCTAGCGGCCAACACTCGCGCAATCGTGTACGCGCTCGTCACCGCACTAGGGATGGACCCCGACCGACTGCTGCTGACGAAAAATAATAATTGCACTTCTACAGACCTCACGACTGATTACGACTGA
>DPOGS212432-PA
MGDFLVMGIKLKIILLCTAVKTVTSTGLGKCPIYPQFPDYDINRMTGTWYEVERSFHLMEISASCIQLDVTINERGFLLITVNTINRWTNTPSISYGLGIPSHNGSSSFRYKLNNRMPYIIGRMLPGAGLYNVLFTDYERFAIIWSCTNYSIAHADRLWILGRHRSLAANTRAIVYALVTALGMDPDRLLLTKNNNCTSTDLTTDYD-