| DPOGS212557 | ||
|---|---|---|
| Transcript | DPOGS212557-TA | 561 bp |
| Protein | DPOGS212557-PA | 186 aa |
| Genomic position | DPSCF300075 - 403201-404557 | |
| RNAseq coverage | 192x (Rank: top 48%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL007773 | 6e-45 | 48.88% | |
| Bombyx | BGIBMGA012280-TA | 8e-74 | 100.00% | |
| Drosophila | CG7646-PA | 7e-104 | 95.16% | |
| EBI UniRef50 | UniRef50_Q9VW66 | 7e-102 | 95.16% | CG7646, isoform A n=26 Tax=Bilateria RepID=Q9VW66_DROME |
| NCBI RefSeq | XP_969439.1 | 8e-103 | 95.70% | PREDICTED: similar to AGAP007248-PA [Tribolium castaneum] |
| NCBI nr blastp | gi|91093205 | 1e-101 | 95.70% | PREDICTED: similar to AGAP007248-PA [Tribolium castaneum] |
| NCBI nr blastx | gi|91093205 | 2e-101 | 95.70% | PREDICTED: similar to AGAP007248-PA [Tribolium castaneum] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005509 | 7.1e-50 | calcium ion binding | |
| KEGG pathway | rno:140936 | 1e-36 | ||
| K13764 (RCVRN) | maps-> | Phototransduction | ||
| InterPro domain | [11-184] IPR011992 | 7.1e-50 | EF-hand-like domain | |
| [10-24] IPR001125 | 3.2e-41 | Recoverin | ||
| [67-91] IPR018248 | 1.4e-07 | EF-hand | ||
| [102-130] IPR002048 | 4.5e-06 | Calcium-binding EF-hand | ||
| Orthology group | MCL16819 | Insect specific | ||
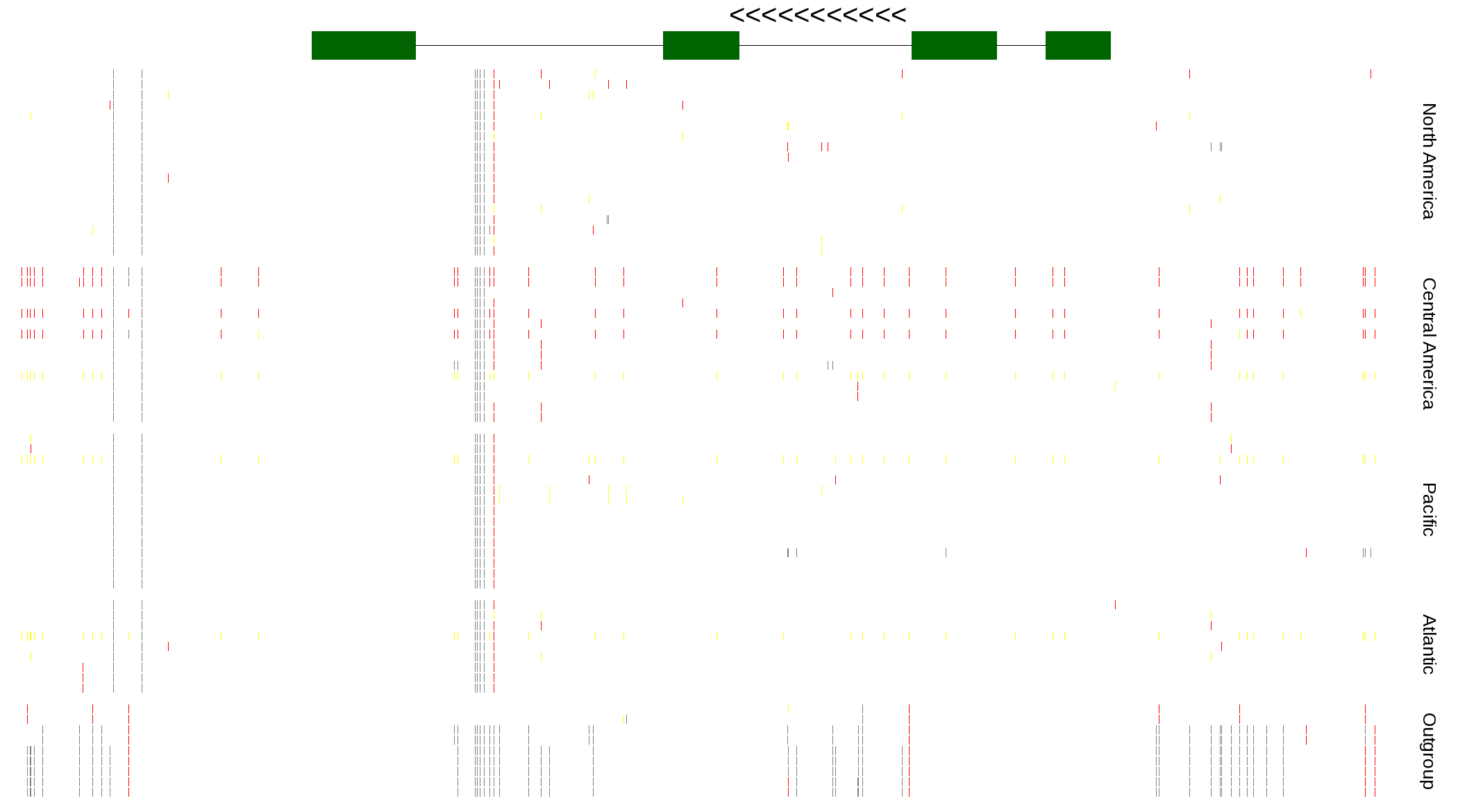
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS212557-TA
ATGGGTTGTTTCGTCAGCAAAGATAAACTTACCAAAGAGGACATGGATTTCCTCAAAACCCATACATCGTACGATGAGACCACCATAAAAGAATGGTACAAGGGTTTCAAGCAAGATTGTCCCAACGGCAGGCTCACGCCGGCTAAATTTGTCGACATGTACAAAATGTTCTTCCCATCTGGTAATGCGGTTGAATTCTGCGACCACGTTTTCCGTACATTCGATATGGACAAGAATGGATACATAGATTTTAAGGAATTCCTGCAAGCCATCCATGTCACCTCTAGCGGGACCCCGGAGGAAAAACTGAAGTGGGCCTTCCGTATGTACGATGTAGATGGGAACGGGGTGATAGATATTCAAGAGATGACGAAAATAGTCCAAGCCATATACGACATGCTTGGCGCTTGCTCCAGCAACAGGCCCGCGGACTCGGCCGAGGAGCGCGCTAAAAACATCTTCGCTAAGATGGACGAGAACAACGACGGTCAATTGACCCAGGACGAGTTCCTCAAAGGCTGCCTCCAGGATGAGGAGCTGTCTAAGATGTTGGCTCCCTAA
>DPOGS212557-PA
MGCFVSKDKLTKEDMDFLKTHTSYDETTIKEWYKGFKQDCPNGRLTPAKFVDMYKMFFPSGNAVEFCDHVFRTFDMDKNGYIDFKEFLQAIHVTSSGTPEEKLKWAFRMYDVDGNGVIDIQEMTKIVQAIYDMLGACSSNRPADSAEERAKNIFAKMDENNDGQLTQDEFLKGCLQDEELSKMLAP-