| DPOGS212600 | ||
|---|---|---|
| Transcript | DPOGS212600-TA | 618 bp |
| Protein | DPOGS212600-PA | 205 aa |
| Genomic position | DPSCF300245 - 168136-168823 | |
| RNAseq coverage | 98x (Rank: top 61%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL002477 | 6e-73 | 68.06% | |
| Bombyx | BGIBMGA008503-TA | 6e-61 | 57.29% | |
| Drosophila | CG11454-PA | 8e-21 | 49.35% | |
| EBI UniRef50 | UniRef50_UPI000224615C | 1e-27 | 36.36% | UPI000224615C related cluster n=1 Tax=unknown RepID=UPI000224615C |
| NCBI RefSeq | XP_307927.4 | 1e-26 | 58.43% | AGAP002256-PA [Anopheles gambiae str. PEST] |
| NCBI nr blastp | gi|345478874 | 4e-27 | 36.36% | PREDICTED: hypothetical protein LOC100123901 [Nasonia vitripennis] |
| NCBI nr blastx | gi|345478874 | 7e-28 | 34.91% | PREDICTED: hypothetical protein LOC100123901 [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003676 | 1.2e-17 | nucleic acid binding | |
| GO:0000166 | 2.3e-16 | nucleotide binding | ||
| KEGG pathway | ||||
| InterPro domain | [8-80] IPR000504 | 1.2e-17 | RNA recognition motif domain | |
| [3-79] IPR012677 | 2.3e-16 | Nucleotide-binding, alpha-beta plait | ||
| Orthology group | MCL19557 | Insect specific | ||
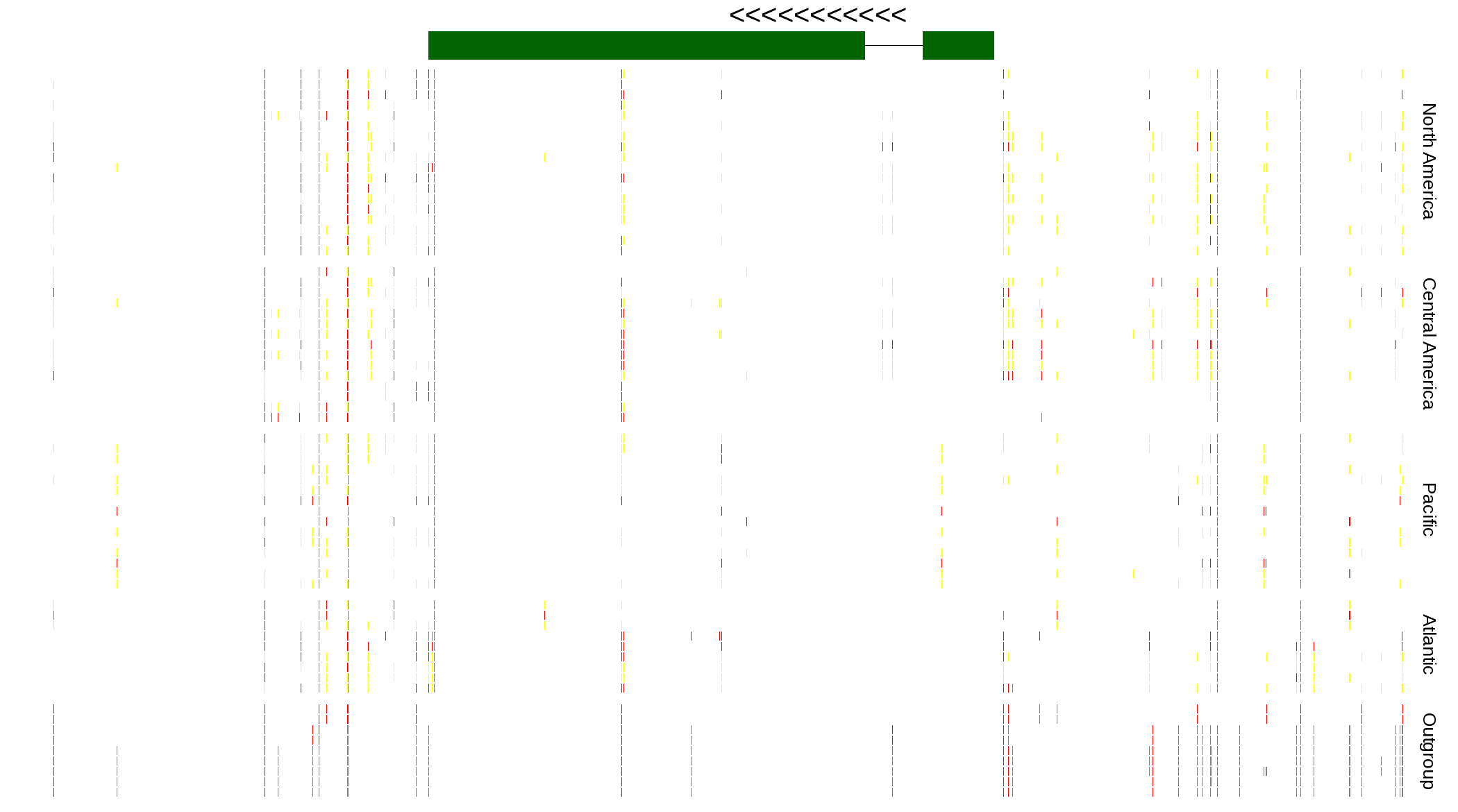
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS212600-TA
ATGATCGAAGAGGCCAACAAGACTGTTTGGTGTGGAAATTTATCGGAACAAGTGACCGAAGAACTTCTTTATGAGTTATTTGTTCAAGCAGGTCCAGTAGAGAAGGTTATAATACCAAAAGACAAAGATGGTAGACAAAAGAATTTCGCTTTTATAACATACTGCCATGAAGTATCCGTGCCATATGCCATTAACCTATTCCGTGGCACAGCACTGTTTCATAGAACTCTGCTTCTACAGAATAGAGGTCGAATGGCGTTATTGCCACCACCAATTAGATGCTATGGCCCTGAAGCCAGTCTCGACTTTAATGCTCAATCGAGTGTAACACAACAGTTCACTGATATGACTGATAAGCTTAAGGAGGAAAATCCCCAACCAGCCCACTTACCCACACAGAGACAAGACATGAGCGACAAACTTGTAGTTGCTAGTTTGCAGGGTAATTGGTCTAACCGACACCATCCATATAGGTCTAAAGAGAAATTTCATAATCGCGACGACTACAGACCTAGAGACGGCCATGACAATTATAACAGAGTTAAAACAAGCCATTGGAGAGAAAGACGGGCCCAATCCTGCTCCTCCGCGCTAATTGAATTAGGATCTAACATCTAA
>DPOGS212600-PA
MIEEANKTVWCGNLSEQVTEELLYELFVQAGPVEKVIIPKDKDGRQKNFAFITYCHEVSVPYAINLFRGTALFHRTLLLQNRGRMALLPPPIRCYGPEASLDFNAQSSVTQQFTDMTDKLKEENPQPAHLPTQRQDMSDKLVVASLQGNWSNRHHPYRSKEKFHNRDDYRPRDGHDNYNRVKTSHWRERRAQSCSSALIELGSNI-