| DPOGS213177 | ||
|---|---|---|
| Transcript | DPOGS213177-TA | 315 bp |
| Protein | DPOGS213177-PA | 104 aa |
| Genomic position | DPSCF300114 - 210795-211369 | |
| RNAseq coverage | 475x (Rank: top 26%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL010298 | 4e-49 | 78.85% | |
| Bombyx | BGIBMGA007367-TA | 4e-48 | 75.00% | |
| Drosophila | nmdyn-D6-PA | 2e-19 | 53.42% | |
| EBI UniRef50 | UniRef50_E1ZZR2 | 1e-24 | 52.48% | Nucleoside diphosphate kinase 6 n=5 Tax=Pancrustacea RepID=E1ZZR2_CAMFO |
| NCBI RefSeq | XP_001602836.1 | 3e-29 | 55.45% | PREDICTED: hypothetical protein [Nasonia vitripennis] |
| NCBI nr blastp | gi|156548316 | 7e-28 | 55.45% | PREDICTED: nucleoside diphosphate kinase 6-like [Nasonia vitripennis] |
| NCBI nr blastx | gi|156548316 | 1e-26 | 55.45% | PREDICTED: nucleoside diphosphate kinase 6-like [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0006228 | 3.9e-24 | UTP biosynthetic process | |
| GO:0005524 | 3.9e-24 | ATP binding | ||
| GO:0006241 | 3.9e-24 | CTP biosynthetic process | ||
| GO:0006165 | 3.9e-24 | nucleoside diphosphate phosphorylation | ||
| GO:0004550 | 3.9e-24 | nucleoside diphosphate kinase activity | ||
| GO:0006183 | 3.9e-24 | GTP biosynthetic process | ||
| KEGG pathway | nvi:100118978 | 9e-29 | ||
| K00940 (E2.7.4.6, ndk) | maps-> | Purine metabolism | ||
| Pyrimidine metabolism | ||||
| InterPro domain | [15-74] IPR001564 | 3.9e-24 | Nucleoside diphosphate kinase | |
| Orthology group | MCL13193 | Single-copy universal gene | ||
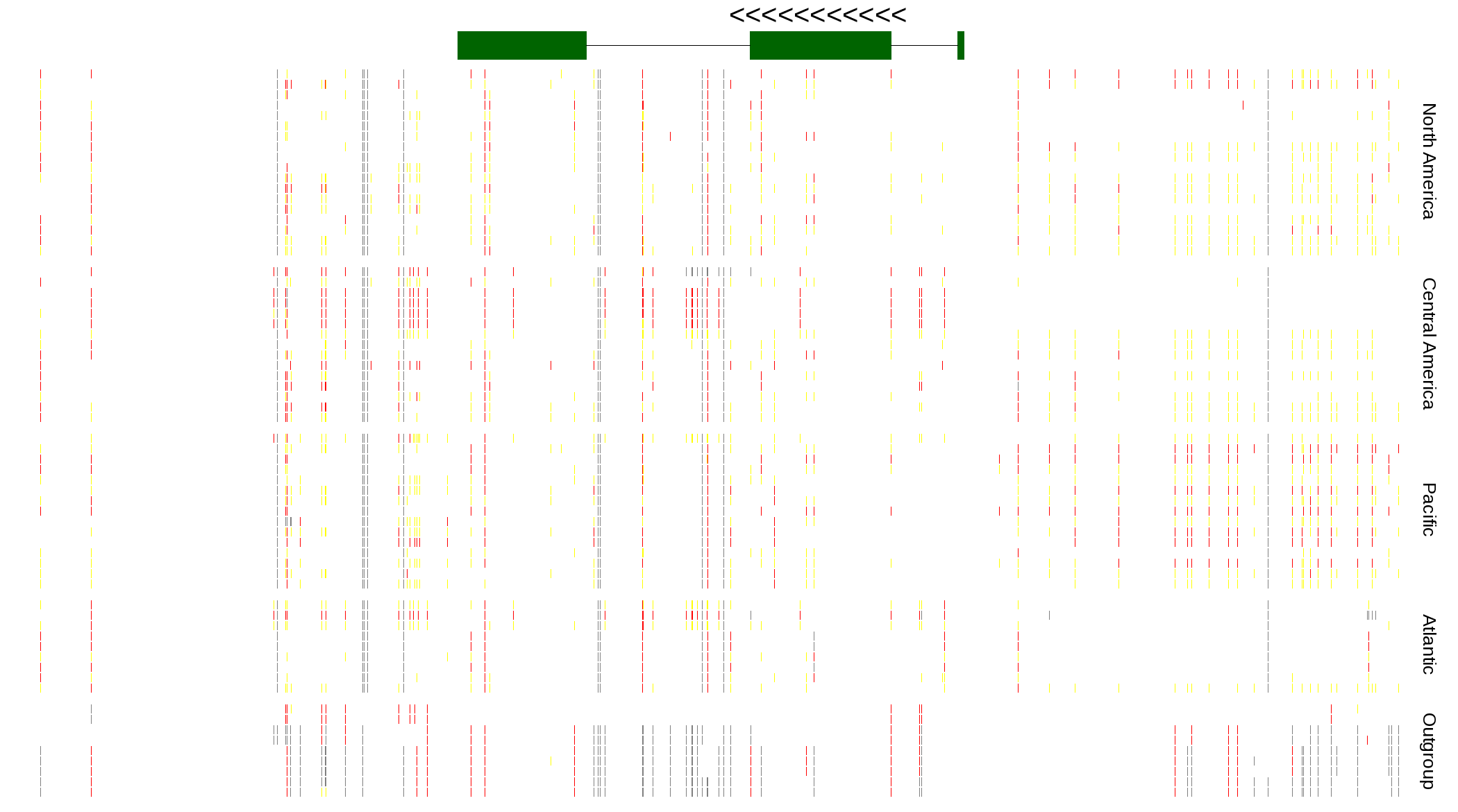
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS213177-TA
ATGAGCAGCGGTTGCGTGGATCTCCACATCATTGGACACACCAATGCTATCGAACTATGGAGGAAAATGCTCGGTCCAACCAAAGTATATAGAGCGCAGTATCAAGAGCCCTACTGTTTGAGAGGCATGTTCGGATTGTCTGATACTAGGAATGTCGCTCATGGTTCCGATTCAGAAACATCCGCTGAGAGGGAAATCAAATTCTTCTTCCCAGATTTCTCATTCTATAAGTGGCATACCTCGGATGAAATGACTTTCCGGAAAGGACCAATAATATTCAATCACCACATGTTCCAGCACGTCAGGAGGCTGTAA
>DPOGS213177-PA
MSSGCVDLHIIGHTNAIELWRKMLGPTKVYRAQYQEPYCLRGMFGLSDTRNVAHGSDSETSAEREIKFFFPDFSFYKWHTSDEMTFRKGPIIFNHHMFQHVRRL-