| DPOGS213829 | ||
|---|---|---|
| Transcript | DPOGS213829-TA | 294 bp |
| Protein | DPOGS213829-PA | 97 aa |
| Genomic position | DPSCF300183 - 146457-151730 | |
| RNAseq coverage | 232x (Rank: top 44%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL012684 | 4e-27 | 56.99% | |
| Bombyx | BGIBMGA005429-TA | 5e-14 | 58.62% | |
| Drosophila | Acyp2-PA | 1e-23 | 46.15% | |
| EBI UniRef50 | UniRef50_G3WHR7 | 1e-26 | 56.99% | Acylphosphatase n=6 Tax=Coelomata RepID=G3WHR7_SARHA |
| NCBI RefSeq | NP_001155440.1 | 4e-28 | 57.45% | hypothetical protein LOC100160410 [Acyrthosiphon pisum] |
| NCBI nr blastp | gi|383862697 | 2e-27 | 61.11% | PREDICTED: acylphosphatase-1-like [Megachile rotundata] |
| NCBI nr blastx | gi|383862697 | 9e-28 | 61.11% | PREDICTED: acylphosphatase-1-like [Megachile rotundata] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0003998 | 4.3e-26 | acylphosphatase activity | |
| KEGG pathway | api:100160410 | 1e-27 | ||
| K01512 (E3.6.1.7, acyP) | maps-> | Benzoate degradation via CoA ligation | ||
| Pyruvate metabolism | ||||
| InterPro domain | [1-95] IPR001792 | 1.2e-29 | Acylphosphatase-like | |
| [5-20] IPR020456 | 4.3e-26 | Acylphosphatase | ||
| Orthology group | MCL11605 | Multiple-copy universal gene | ||
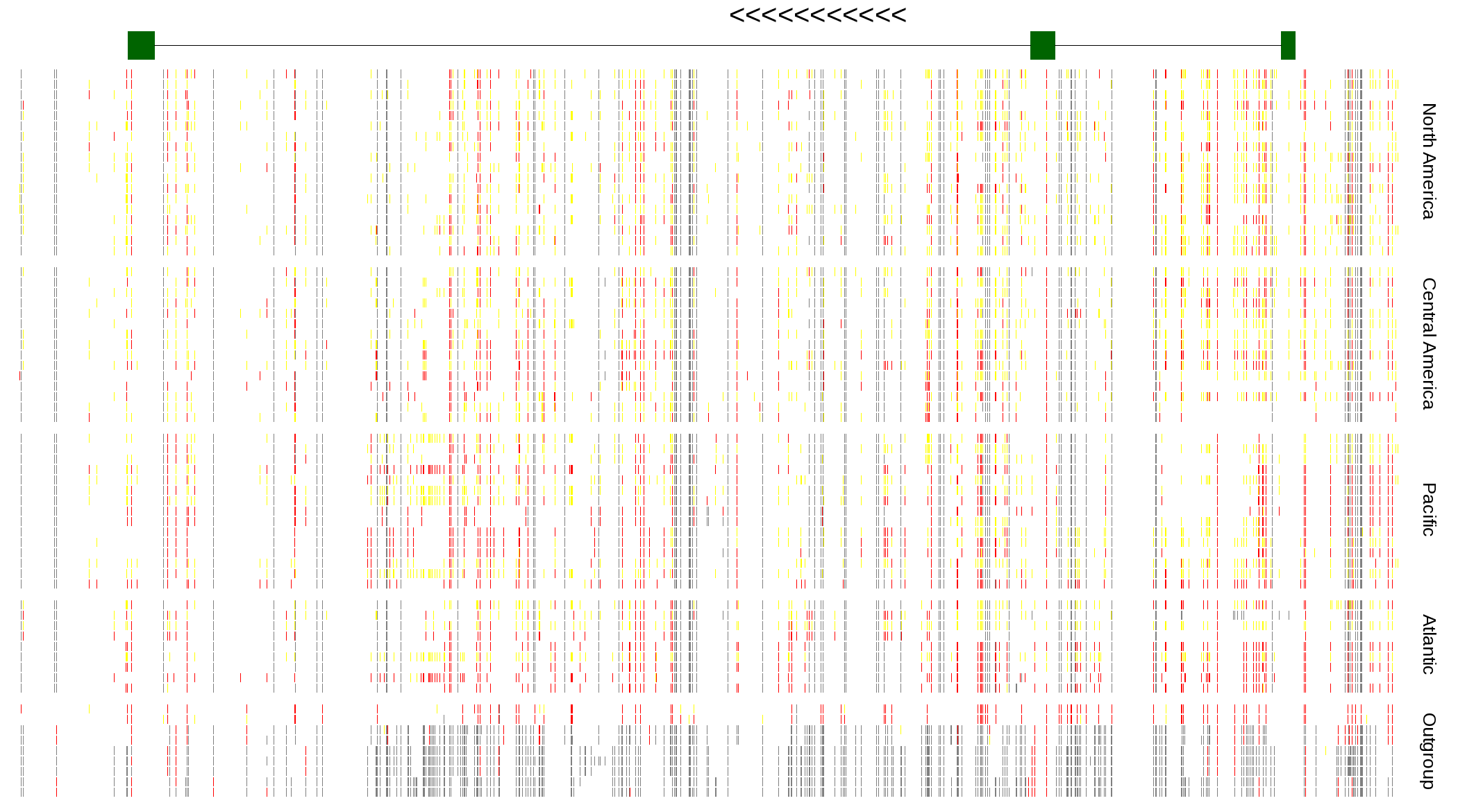
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS213829-TA
ATGGCACTCCGAAGCGTAGACTTTGAAGTGTATGGCCGGGTTCAAGGCGTATTCTTTAGAAAGTTCACAAAAGAGCAAGCAGAGAAATTGGCACTGAAAGGCTGGTGTCGTAATACTTCTCGCGGGACTGTCGAGGGACAGGTTCAGGGTCCGTCGGAAAAAGTCGATGAAATGATGCATTGGCTAAAAACCACGGGAAGTCCGAGTAGTAAGATTGAAAAAGCAGATTTCAGAAACGGCAAAGATATATCCGAATACAATTTTAGCAATTTCGAAATACGACGCGATGATTAG
>DPOGS213829-PA
MALRSVDFEVYGRVQGVFFRKFTKEQAEKLALKGWCRNTSRGTVEGQVQGPSEKVDEMMHWLKTTGSPSSKIEKADFRNGKDISEYNFSNFEIRRDD-