| DPOGS214025 | ||
|---|---|---|
| Transcript | DPOGS214025-TA | 507 bp |
| Protein | DPOGS214025-PA | 168 aa |
| Genomic position | DPSCF300238 - 220272-221098 | |
| RNAseq coverage | 0x (Rank: top 99%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL011168 | 3e-14 | 34.59% | |
| Bombyx | BGIBMGA009715-TA | 2e-09 | 41.89% | |
| Drosophila | % | |||
| EBI UniRef50 | UniRef50_P43515 | 3e-10 | 33.99% | Chorion class B protein Ld10 n=4 Tax=Lymantria dispar RepID=CHB1_LYMDI |
| NCBI RefSeq | NP_001112382.1 | 7e-06 | 42.03% | chorion class B protein ERB4 precursor [Bombyx mori] |
| NCBI nr blastp | gi|1168928 | 9e-10 | 33.99% | chorion protein [Lymantria dispar] |
| NCBI nr blastx | gi|1168928 | 5e-11 | 33.33% | chorion protein [Lymantria dispar] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0007304 | 1.6e-18 | chorion-containing eggshell formation | |
| GO:0005213 | 1.6e-18 | structural constituent of chorion | ||
| GO:0007275 | 1.6e-18 | multicellular organismal development | ||
| GO:0042600 | 1.6e-18 | chorion | ||
| KEGG pathway | ||||
| InterPro domain | [3-148] IPR002635 | 1.6e-18 | Chorion protein | |
| Orthology group | MCL27841 | Specific divergent | ||
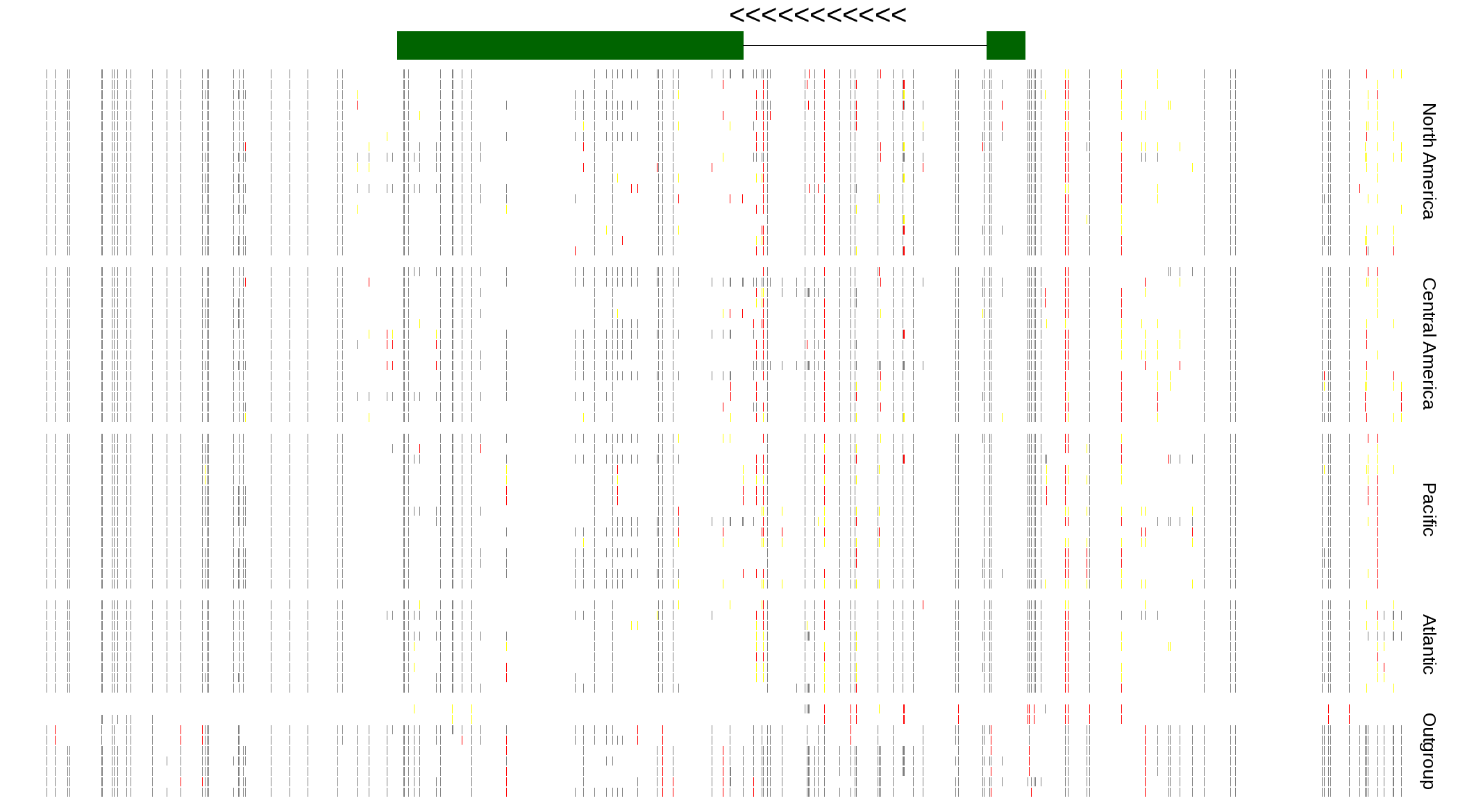
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS214025-TA
ATGGCAGCCAAAACTACCCTCTTTTTCTGCGCTGCGGCGCTTTTAATCCAGGACATATTCGGGGTTTATATCGATCCGTCATATGATAGAATTGCTCCCTGGGCTAGGACATTCTTAGCTTGGAGGGAAGCAAATACAAATGTCGCTGGTTCTTCGAAGCCACTTCTTAAAGTACCTAGGGCTTGTCAATACTCTTATGATGTACCAGTTTATGGTACGCCATACGGAGCTTATAGCGAGATCGCCCTGCCATCTGATTTTCCCACTGATGCTTTGATCGTTACCAGTTCTTCAAATGGACCTATTGGAGTTTCAATACTATCAGAGAATGTAATAAAAGGACTGTTAGCTGTTGGAGGAGAGCTCCCTTTCTTGGGTTCTGTCAGTTTAGAAGGAGTTTTGCCGACTACAGGGTCTGGTACAATTTCCTACAGTTCTGGGGCCAACAGACATGGCCCTGTTATCTGTAAAGAATTAAATCGTCCTGGCAGTCCACTGCCATACTGA
>DPOGS214025-PA
MAAKTTLFFCAAALLIQDIFGVYIDPSYDRIAPWARTFLAWREANTNVAGSSKPLLKVPRACQYSYDVPVYGTPYGAYSEIALPSDFPTDALIVTSSSNGPIGVSILSENVIKGLLAVGGELPFLGSVSLEGVLPTTGSGTISYSSGANRHGPVICKELNRPGSPLPY-