| DPOGS215107 | ||
|---|---|---|
| Transcript | DPOGS215107-TA | 528 bp |
| Protein | DPOGS215107-PA | 175 aa |
| Genomic position | DPSCF300139 - 8614-10077 | |
| RNAseq coverage | 152x (Rank: top 53%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL004579 | 1e-74 | 69.71% | |
| Bombyx | BGIBMGA009598-TA | 8e-61 | 59.43% | |
| Drosophila | Egm-PA | 3e-28 | 37.87% | |
| EBI UniRef50 | UniRef50_G3MRD7 | 2e-27 | 48.12% | Putative uncharacterized protein n=1 Tax=Amblyomma maculatum RepID=G3MRD7_9ACAR |
| NCBI RefSeq | XP_002033416.1 | 3e-27 | 38.92% | GM20419 [Drosophila sechellia] |
| NCBI nr blastp | gi|321470919 | 2e-27 | 39.39% | hypothetical protein DAPPUDRAFT_302851 [Daphnia pulex] |
| NCBI nr blastx | gi|321470919 | 5e-27 | 39.39% | hypothetical protein DAPPUDRAFT_302851 [Daphnia pulex] |
| Group | ||||
|---|---|---|---|---|
| KEGG pathway | xtr:548505 | 8e-10 | ||
| K00257 (E1.3.99.-) | maps-> | Geraniol degradation | ||
| 1- and 2-Methylnaphthalene degradation | ||||
| Orthology group | ||||
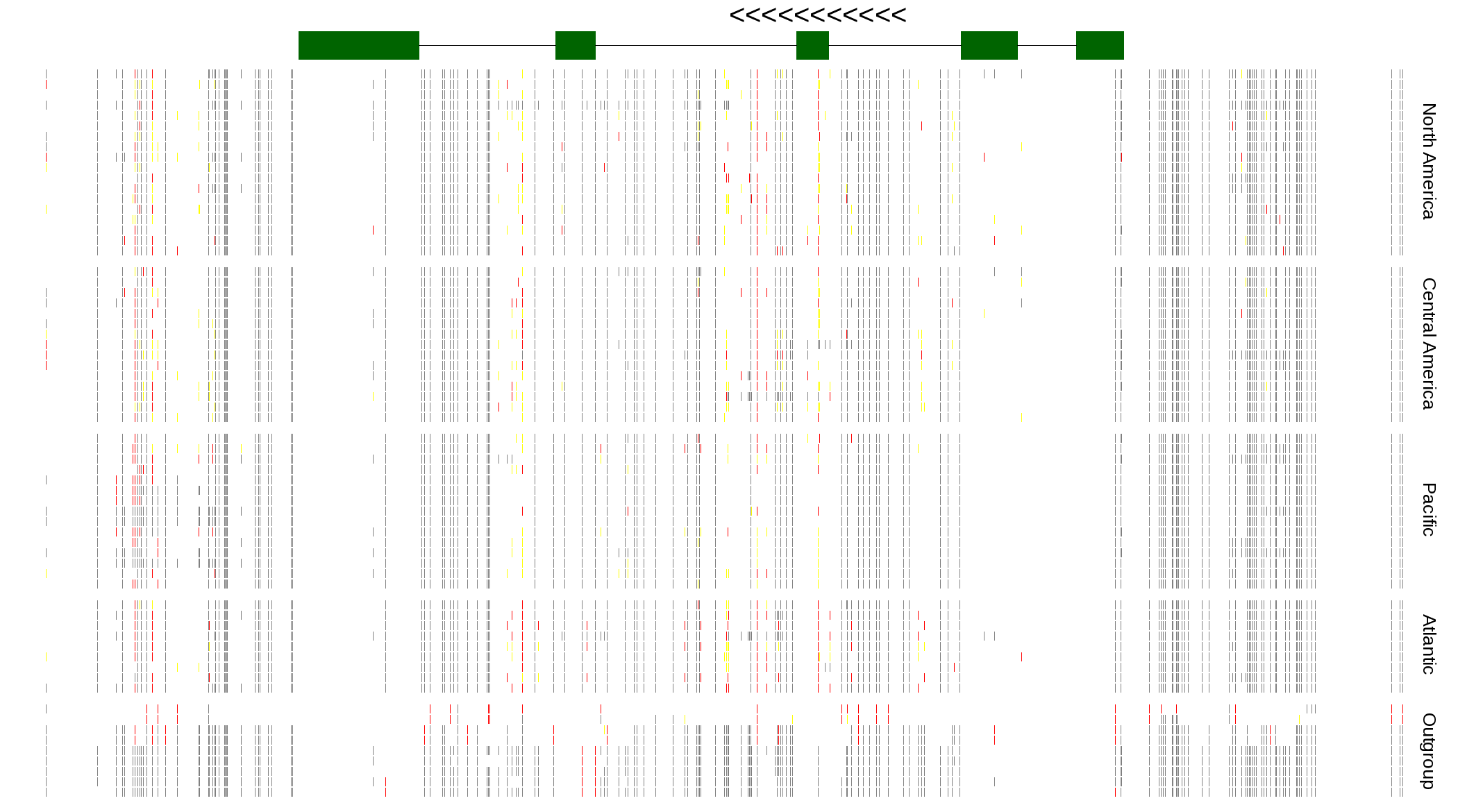
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS215107-TA
ATGGCGACGGACGTGAAAAGGCTAAGGAATCCGCTATTCAACCCATCGTTCATCATAAAGAAGATCATCGCTGACAGAAACCAGGAGAAGGACGAACCTAGACTGGACCTGTACCTATCGGAGGACCTCCATCCAACTTTGAAGAAACCCTCGGAACAGCTGGAGTACTGCGTTCTTAGGATGAGGTACATAGTGGAATCTTTAATGAGCAGATATGGAACGGACGTGGTGATGGCCGGCACCGAGTTAAACCGGTTGGCTGAAGCATCTACGCAGATATATGTCATGGCTGCAGTACTGGCGAGGGCTTCCAGGGCATATTGCATAGGATTGAGGAACGGTGAGTTTGAAATGAAAATTGCAGCTTGTTTTGTTGAAAGCACCAAAGATCATGTTAAGAAACTGCTCTTGGAGATAAGCGAAGGGGAGTACCTTAATATGGACCACTTCAGGATAAATTTCGGCAAGACCGTCCTCGATAATAAGGGCCAGTTGCTTGAAAAGCCAACCGCGAGGGTTTTTTGGTAG
>DPOGS215107-PA
MATDVKRLRNPLFNPSFIIKKIIADRNQEKDEPRLDLYLSEDLHPTLKKPSEQLEYCVLRMRYIVESLMSRYGTDVVMAGTELNRLAEASTQIYVMAAVLARASRAYCIGLRNGEFEMKIAACFVESTKDHVKKLLLEISEGEYLNMDHFRINFGKTVLDNKGQLLEKPTARVFW-