| DPOGS215353 | ||
|---|---|---|
| Transcript | DPOGS215353-TA | 606 bp |
| Protein | DPOGS215353-PA | 201 aa |
| Genomic position | DPSCF300351 - 103627-113319 | |
| RNAseq coverage | 146x (Rank: top 54%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL005226 | 1e-79 | 85.96% | |
| Bombyx | BGIBMGA009561-TA | 3e-54 | 94.44% | |
| Drosophila | CG5890-PB | 3e-79 | 67.12% | |
| EBI UniRef50 | UniRef50_Q177Q6 | 2e-80 | 70.62% | Potassium channel interacting protein n=3 Tax=Endopterygota RepID=Q177Q6_AEDAE |
| NCBI RefSeq | XP_001868842.1 | 6e-82 | 72.04% | potassium channel interacting protein [Culex quinquefasciatus] |
| NCBI nr blastp | gi|170068375 | 1e-80 | 72.04% | potassium channel interacting protein [Culex quinquefasciatus] |
| NCBI nr blastx | gi|170068375 | 4e-79 | 72.04% | potassium channel interacting protein [Culex quinquefasciatus] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005509 | 1.4e-31 | calcium ion binding | |
| KEGG pathway | cfa:489500 | 1e-20 | ||
| K13764 (RCVRN) | maps-> | Phototransduction | ||
| InterPro domain | [29-199] IPR011992 | 1.4e-31 | EF-hand-like domain | |
| [24-38] IPR001125 | 2.3e-11 | Recoverin | ||
| [107-131] IPR018248 | 1e-06 | EF-hand | ||
| Orthology group | MCL15919 | Insect specific | ||
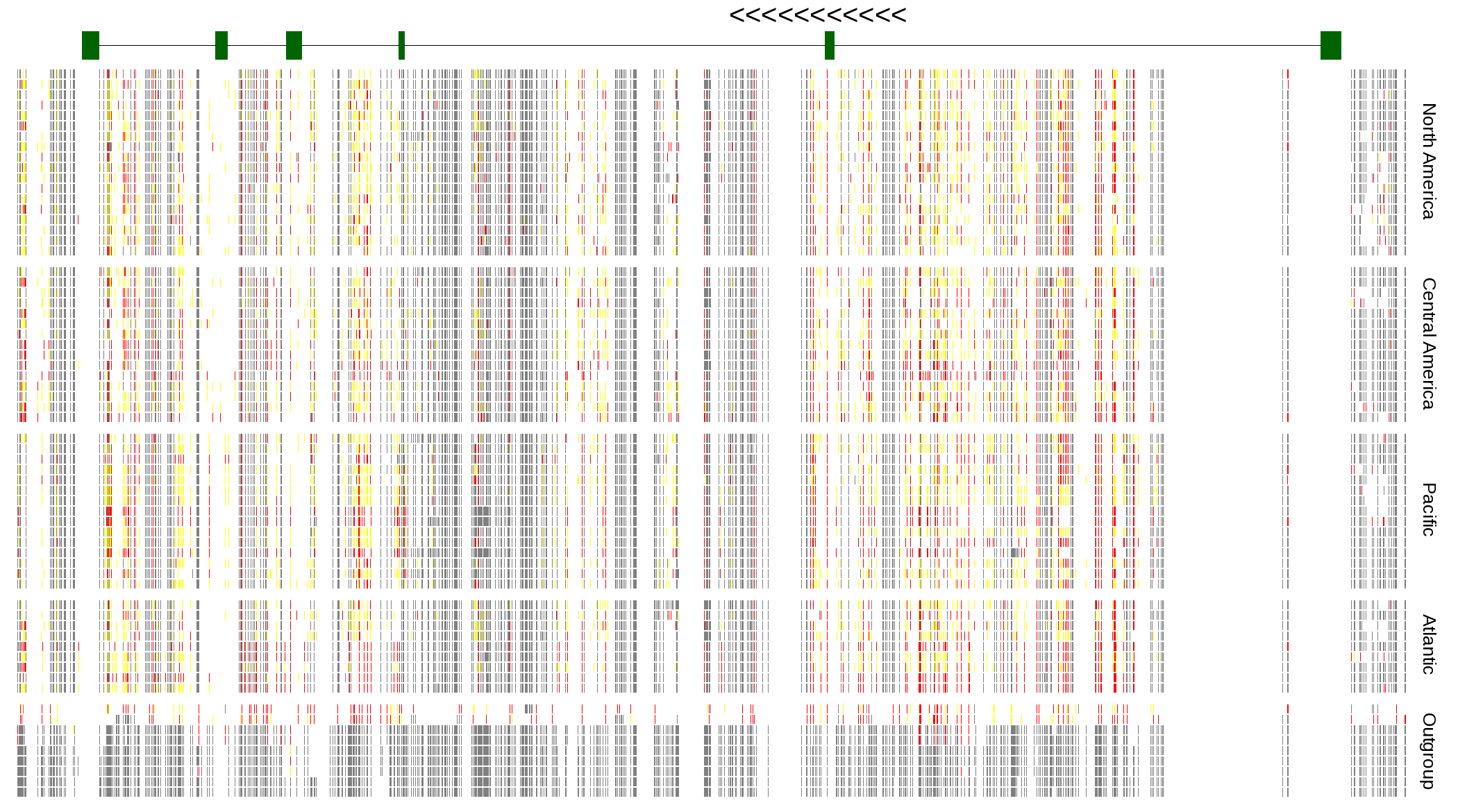
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS215353-TA
ATGGCTACGCCACCGGAGTCTCCCGTGGAGGAGCACGCATACGAGCTTGAACCGAACCGTGCACCCAAACCAATACCAGTGGCTCTCGAGGAGTTGTGCAGACAAACCAAGTTCTCCAAACAGGAGATCAGGGTTATGTACAGGGGGTTCAAGACGGAATGTCCGGAAGGGGTAGTCCACGAGGATTCTTTCAAAGACATTTACGCCAAGTTCTTCCCACACGGAATTTTGCATGCGCGTTCAGTTCAGAATGACAGCTGTTTGAGGGAACTATTGGTGACCCTGTCGACTCTGCTGCGAGGGTCGGTGTACGAGAGACTGAGGTGGGCATTCAGGTTGTACGACGTGGATGGTGACGGGGCCATCACCAGGCAAGAACTGGCCGAGGTGGTGGTAGCTGTTCACGAGCTGCTGGGTCGCCGGGCGCCGCCGGGCTCACCCGCAGCCAGGATCGATGACGCGAAGGCTAACGAACAGGTGGACCGGGTGTTCAGGAAGCTGGACCTGAACCAGGACGGCGTCATCACCATCGAGGAGTTCCTGGAGTCGTGCCTCAAGGATGACGTCATCACGCGCTCGCTGCAGATGTTCGACACGGTGCTATGA
>DPOGS215353-PA
MATPPESPVEEHAYELEPNRAPKPIPVALEELCRQTKFSKQEIRVMYRGFKTECPEGVVHEDSFKDIYAKFFPHGILHARSVQNDSCLRELLVTLSTLLRGSVYERLRWAFRLYDVDGDGAITRQELAEVVVAVHELLGRRAPPGSPAARIDDAKANEQVDRVFRKLDLNQDGVITIEEFLESCLKDDVITRSLQMFDTVL-