| DPOGS215633 | ||
|---|---|---|
| Transcript | DPOGS215633-TA | 612 bp |
| Protein | DPOGS215633-PA | 203 aa |
| Genomic position | DPSCF300041 - 1858472-1859963 | |
| RNAseq coverage | 118x (Rank: top 58%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006536 | 6e-29 | 82.61% | |
| Bombyx | % | |||
| Drosophila | Gem2-PA | 3e-13 | 31.15% | |
| EBI UniRef50 | UniRef50_E1ZZS8 | 3e-26 | 37.27% | Survival of motor neuron protein-interacting protein 1 n=7 Tax=Formicidae RepID=E1ZZS8_CAMFO |
| NCBI RefSeq | XP_001600147.1 | 1e-25 | 38.83% | PREDICTED: hypothetical protein [Nasonia vitripennis] |
| NCBI nr blastp | gi|332029975 | 5e-30 | 40.28% | Survival of motor neuron protein-interacting protein 1 [Acromyrmex echinatior] |
| NCBI nr blastx | gi|332029975 | 7e-29 | 40.28% | Survival of motor neuron protein-interacting protein 1 [Acromyrmex echinatior] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0000398 | 4.6e-17 | nuclear mRNA splicing, via spliceosome | |
| GO:0000245 | 4.6e-17 | spliceosome assembly | ||
| KEGG pathway | ||||
| InterPro domain | [137-203] IPR007022 | 4.6e-17 | Survival motor neuron interacting protein 1 | |
| Orthology group | MCL15342 | Single-copy universal gene | ||
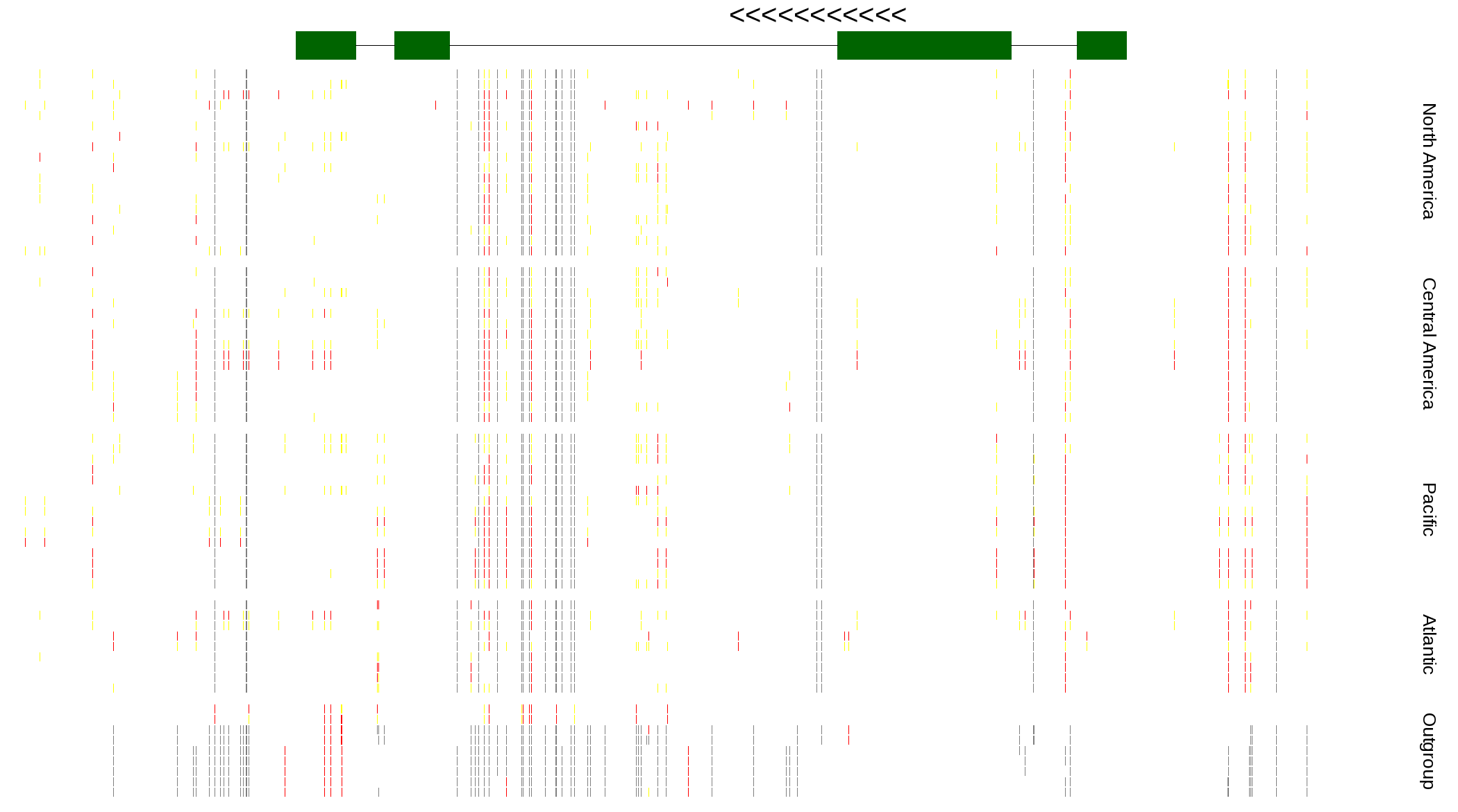
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS215633-TA
ATGAAAGAAAGGCAAAATTATTCAACAGTTACAACTTGTAACAGGGATTTTTCCAAGTTTGCCCGAAACCAATCATGTTTCGTTAAGGAGTTACCACATGCTAAAGCACCAGAGAGTCTAAAACCAACTATAGAATGGCAAAATATTCAAGTTGCTGATTTCTCAAAGGTTAGAATGTACATTTCAAAGTTGATATCTAATAGATCACTTTGGCCTAGAGATGTTATAAATATAGAAATAGATCCAGACAACATTGCAGCTTGGATGAACTTATTTGAGAATAAAGACCCAAAGCTTTCTTGTGTGCTTGGCTTGCATCATGCATTGTTAGACCATGGCCTGGAGATTCTCATTGAAATGCTGGATAAAGTCAAACCTGGCAGTACAATAAACTATAAAACAGGTCAATGGATTTACGCATTTTTAGCCTGCACCAGACAACCTTTGCTCTCTGACACTATAAGTATATTAAGAAACCTAGCTAGAAAATGTGCAGAGATTAGATCTCATCTCAATACAGAGGATGTGAGTTCCAGGGAGGCGGCCGCTCCATTAAACATATTTATATGCTTGGTAGCTCGATACTTCAGACAATATGATCTTGCTGATTGA
>DPOGS215633-PA
MKERQNYSTVTTCNRDFSKFARNQSCFVKELPHAKAPESLKPTIEWQNIQVADFSKVRMYISKLISNRSLWPRDVINIEIDPDNIAAWMNLFENKDPKLSCVLGLHHALLDHGLEILIEMLDKVKPGSTINYKTGQWIYAFLACTRQPLLSDTISILRNLARKCAEIRSHLNTEDVSSREAAAPLNIFICLVARYFRQYDLAD-