| DPOGS200873 | ||
|---|---|---|
| Transcript | DPOGS200873-TA | 669 bp |
| Protein | DPOGS200873-PA | 222 aa |
| Genomic position | DPSCF300071 + 639534-640202 | |
| RNAseq coverage | 2082x (Rank: top 6%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL011478 | 1e-108 | 87.39% | |
| Bombyx | BGIBMGA009864-TA | 4e-106 | 81.70% | |
| Drosophila | CG13887-PB | 5e-71 | 58.77% | |
| EBI UniRef50 | UniRef50_Q9W0M4 | 6e-69 | 58.77% | CG13887, isoform B n=17 Tax=Diptera RepID=Q9W0M4_DROME |
| NCBI RefSeq | XP_002093049.1 | 1e-73 | 57.89% | GE21102 [Drosophila yakuba] |
| NCBI nr blastp | gi|195490225 | 3e-72 | 57.89% | GE21102 [Drosophila yakuba] |
| NCBI nr blastx | gi|156551872 | 3e-70 | 64.19% | PREDICTED: B-cell receptor-associated protein 31-like isoform 1 [Nasonia vitripennis] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0006886 | 8.8e-104 | intracellular protein transport | |
| GO:0005783 | 8.8e-104 | endoplasmic reticulum | ||
| GO:0016021 | 8.8e-104 | integral to membrane | ||
| KEGG pathway | dya:Dyak_GE21102 | 4e-73 | ||
| K14009 (BCAP31, BAP31) | maps-> | Protein processing in endoplasmic reticulum | ||
| InterPro domain | [2-222] IPR008417 | 8.8e-104 | B-cell receptor-associated 31-like | |
| Orthology group | MCL13905 | Single-copy universal gene | ||
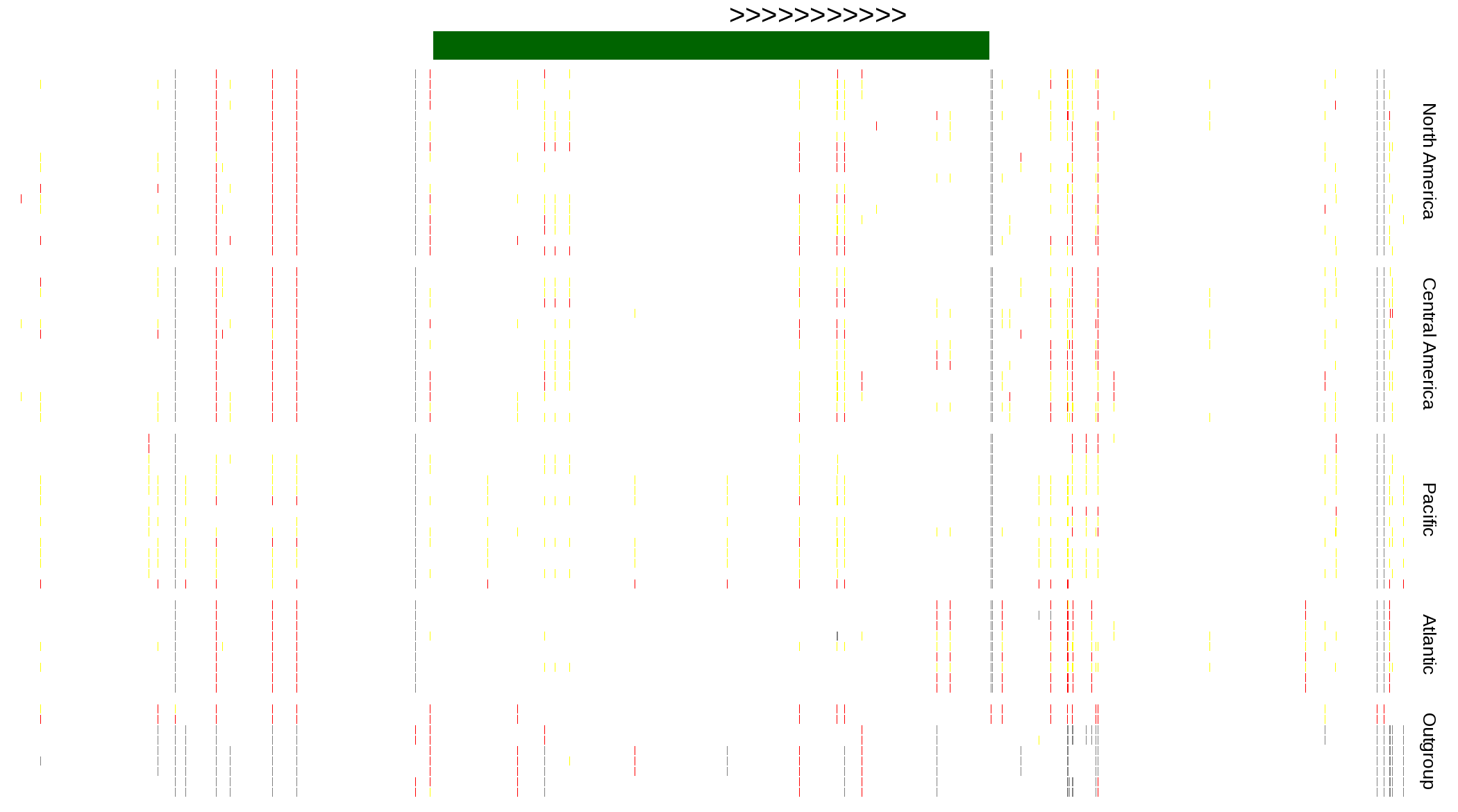
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS200873-TA
ATGAGTCTTCAGTGGACCATCATCGCGTCATTTCTATACGCCGAAATAGCATTTGTACTGTTATTGACCTTACCGATCGCGAGTCCTGCAAGATGGAACAAGTTCTTCAAATCGAAGTTTCTAGCTTACATGACAGGCCAAGCATCCATCTACTTTGTTGTTCTCATTGGGGTTCTAGTTCTCTGCCTTCTCGACGCCATTCGTGAGATTCAGAAGTATTCAAACGTTGAATCTTCAGACCACCAACATTTGGATGCTGAGATGCAAGGAAACATGCGTCTGTTTAGAGCTCAAAGGAACTTGTACATATCAGGTATTGCACTTTTCCTTCTAGTCGTTATTCGGCGCTTGATCCAGATGATCTGTGAGCTGGCGACCTTGTACGCCCAATCCGAAGCGAACTTCCGTCAGGCTCAAAGCGCGTCTGTAGCCGCTAAAGCCCTTTTAGAAAAGCAAGGTGCTGGTGATGAGGTCAACAAGAAGGAGATGGAAGATCTCAAGAGCCAGTTATCAGCCTTGGAAAAGGAGTTAGCCAAGGAGAAGAAAGATAAAGAAGCCGTGAAGTCCCAGGCGGAGAGCCTTAACCGTGAATATGATAGACTTGCGGAGGAACACAGCAGGCTACAGAAGAAAATCACAGTTGCTGGTGGTGATAAGAAGGATGAGTAG
>DPOGS200873-PA
MSLQWTIIASFLYAEIAFVLLLTLPIASPARWNKFFKSKFLAYMTGQASIYFVVLIGVLVLCLLDAIREIQKYSNVESSDHQHLDAEMQGNMRLFRAQRNLYISGIALFLLVVIRRLIQMICELATLYAQSEANFRQAQSASVAAKALLEKQGAGDEVNKKEMEDLKSQLSALEKELAKEKKDKEAVKSQAESLNREYDRLAEEHSRLQKKITVAGGDKKDE-