| DPOGS201860 | ||
|---|---|---|
| Transcript | DPOGS201860-TA | 573 bp |
| Protein | DPOGS201860-PA | 190 aa |
| Genomic position | DPSCF300191 - 44633-46164 | |
| RNAseq coverage | 832x (Rank: top 15%) | |
| Annotation | ||||
|---|---|---|---|---|
| Heliconius | HMEL006992 | 3e-58 | 73.30% | |
| Bombyx | BGIBMGA006109-TA | 2e-72 | 78.01% | |
| Drosophila | HP1b-PB | 2e-40 | 50.00% | |
| EBI UniRef50 | UniRef50_G7YNK4 | 7e-43 | 48.10% | Chromobox protein 1 n=3 Tax=Digenea RepID=G7YNK4_CLOSI |
| NCBI RefSeq | NP_001040539.1 | 6e-70 | 77.49% | chromobox-like protein 5 [Bombyx mori] |
| NCBI nr blastp | gi|114052857 | 1e-68 | 77.49% | chromobox-like protein 5 [Bombyx mori] |
| NCBI nr blastx | gi|114052857 | 2e-80 | 77.66% | chromobox-like protein 5 [Bombyx mori] |
| Group | ||||
|---|---|---|---|---|
| Gene Ontology | GO:0005634 | 1.2e-29 | nucleus | |
| KEGG pathway | ||||
| InterPro domain | [124-186] IPR018125 | 1.1e-29 | Chromo shadow, subgroup | |
| [128-185] IPR008251 | 1.2e-29 | Chromo shadow | ||
| [121-188] IPR016197 | 9.3e-24 | Chromo domain-like | ||
| [18-66] IPR023780 | 1.1e-17 | Chromo domain | ||
| [17-69] IPR000953 | 2.6e-17 | Chromo domain/shadow | ||
| [15-23] IPR017984 | 7.9e-09 | Chromo domain subgroup | ||
| Orthology group | MCL25209 | Lepidoptera specific | ||
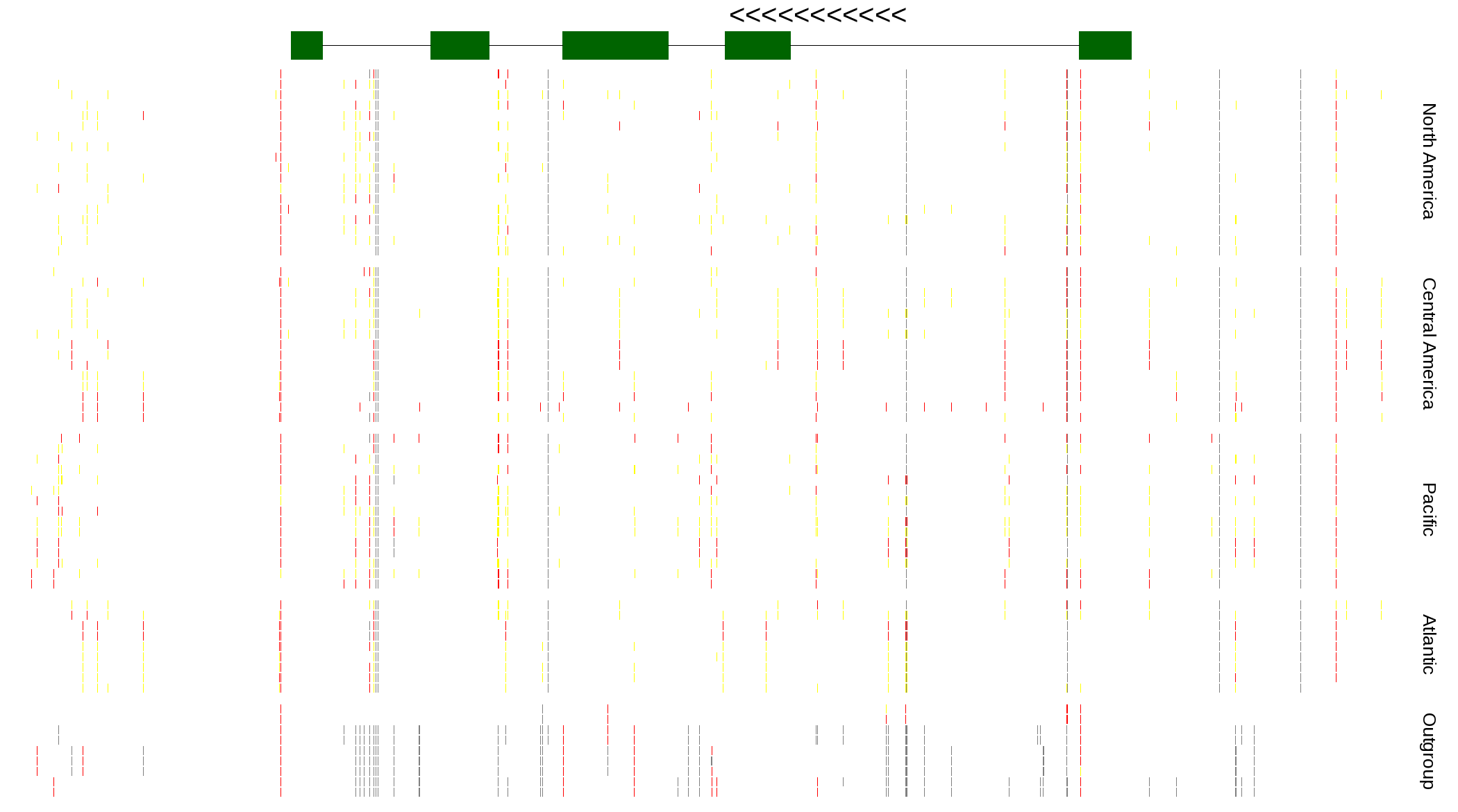
| Genotypes for resequenced monarchs and outgroup Danaus species |
|---|

>DPOGS201860-TA
ATGGGCAAAGATAAAAAGGATGAGACCAGTGAGGGTTCAAGTGAAGAAGAATATGTTGTTGAAAAAGTCTTGAATAAAAGAACTGTCAAGGGGAAGGTCCAATATCTTTTAAAGTGGAAGGGCTACAAAGAAGAGGAAAGCACTTGGGAACCGGAGGAACATTTAGACTGTGAAGAACTCATAAAGGCCTTTGAAAATAGCAGGAAAGAAAAAGAGGTGAAGGCAAAAAAATCGGAGGAACGAAGTAAAAAGCGTCGTCGTGATTCAAGTGACGATACGAGCACAACGGGAAAGATACGTGAGGCCAGTGTGTCTTCGGTTGAGGAACAGAAAGAGGCACGCAAGGAGAAGAAAGATGATAAAAATAAGTCTGGCTTTGAAAAAGGCTTGAAAGCTGAGAAAATTATCGGAGCATCTGATGCCACCGGTGAACTAATGTTTTTAATAAAATGGGCCGACTCTGACGAAGCTGAATTAGTCCCTTCAAAAGTTGCTAACATTAAGTGTCCCCAACAGGTTATAGCATTTTATGAAGAGCGACTGACATGGCACAGTCCAGCCGAGTCGGAGTGA
>DPOGS201860-PA
MGKDKKDETSEGSSEEEYVVEKVLNKRTVKGKVQYLLKWKGYKEEESTWEPEEHLDCEELIKAFENSRKEKEVKAKKSEERSKKRRRDSSDDTSTTGKIREASVSSVEEQKEARKEKKDDKNKSGFEKGLKAEKIIGASDATGELMFLIKWADSDEAELVPSKVANIKCPQQVIAFYEERLTWHSPAESE-